Rotaviruses in Wild Ungulates from Germany, 2019-2022
- PMID: 36985140
- PMCID: PMC10058221
- DOI: 10.3390/microorganisms11030566
Rotaviruses in Wild Ungulates from Germany, 2019-2022
Abstract
Rotavirus A (RVA) is an important cause of diarrhea in humans and animals. However, RVA in wild animals has only scarcely been investigated so far. Here, the presence of RVA in wild ungulates hunted between 2019 and 2022 in Brandenburg, Germany, was investigated using real-time RT-PCR and sequencing of RT-PCR products. By analyzing intestinal contents, RVA-RNA was detected in 1.0% (2/197) of wild boar (Sus scrofa), 1.3% (2/152) of roe deer (Capreolus capreolus), and 2.1% (2/95) of fallow deer (Dama dama) but not in 28 red deer (Cervus elaphus) samples. Genotyping identified G3P[13] strains in wild boar, which were closely related to previously described pig and wild boar strains. Genotype G10P[15] strains, closely related to strains from roe deer, sheep, or cattle, were found in roe deer. The strains of fallow deer represented genotype G3P[3], clustering in a group containing different strains from several hosts. The results indicated a low prevalence of RVA in wild ungulates in Germany. Associations of specific genotypes with certain ungulate species seem to exist but should be confirmed by analyses of more samples in the future.
Keywords: detection rate; fallow deer; genotyping; roe deer; rotavirus A; wild boar.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures

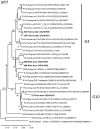

References
-
- Troeger C., Khalil I.A., Rao P.C., Cao S., Blacker B.F., Ahmed T., Armah G., Bines J.E., Brewer T.G., Colombara D.V., et al. Rotavirus Vaccination and the Global Burden of Rotavirus Diarrhea Among Children Younger Than 5 Years. JAMA Pediatr. 2018;172:958–965. doi: 10.1001/jamapediatrics.2018.1960. - DOI - PMC - PubMed
-
- Otto P.H., Rosenhain S., Elschner M.C., Hotzel H., Machnowska P., Trojnar E., Hoffmann K., Johne R. Detection of rotavirus species A, B and C in domestic mammalian animals with diarrhoea and genotyping of bovine species A rotavirus strains. Vet. Microbiol. 2015;179:168–176. doi: 10.1016/j.vetmic.2015.07.021. - DOI - PubMed
-
- Simsek C., Corman V.M., Everling H.U., Lukashev A.N., Rasche A., Maganga G.D., Binger T., Jansen D., Beller L., Deboutte W., et al. At Least Seven Distinct Rotavirus Genotype Constellations in Bats with Evidence of Reassortment and Zoonotic Transmissions. mBio. 2021;12:e02755-20. doi: 10.1128/mBio.02755-20. - DOI - PMC - PubMed
-
- Sachsenröder J., Braun A., Machnowska P., Ng T.F.F., Deng X., Guenther S., Bernstein S., Ulrich R.G., Delwart E., Johne R. Metagenomic identification of novel enteric viruses in urban wild rats and genome characterization of a group A rotavirus. Pt 12J. Gen. Virol. 2014;95:2734–2747. doi: 10.1099/vir.0.070029-0. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

