Total Synthesis and Structure Assignment of the Relacidine Lipopeptide Antibiotics and Preparation of Analogues with Enhanced Stability
- PMID: 37000899
- PMCID: PMC10111413
- DOI: 10.1021/acsinfecdis.3c00043
Total Synthesis and Structure Assignment of the Relacidine Lipopeptide Antibiotics and Preparation of Analogues with Enhanced Stability
Abstract
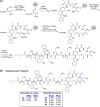
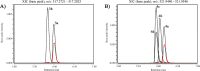
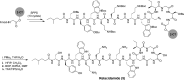
The unabated rise of antibiotic resistance has raised the specter of a post-antibiotic era and underscored the importance of developing new classes of antibiotics. The relacidines are a recently discovered group of nonribosomal lipopeptide antibiotics that show promising activity against Gram-negative pathogens and share structural similarities with brevicidine and laterocidine. While the first reports of the relacidines indicated that they possess a C-terminal five-amino acid macrolactone, an N-terminal lipid tail, and an overall positive charge, no stereochemical configuration was assigned, thereby precluding a full structure determination. To address this issue, we here report a bioinformatics guided total synthesis of relacidine A and B and show that the authentic natural products match our predicted and synthesized structures. Following on this, we also synthesized an analogue of relacidine A wherein the ester linkage of the macrolactone was replaced by the corresponding amide. This analogue was found to possess enhanced hydrolytic stability while maintaining the antibacterial activity of the natural product in both in vitro and in vivo efficacy studies.
Keywords: AMR; NRPS; bioinformatics; lipopeptides; relacidine; total synthesis.
Conflict of interest statement
The authors declare the following competing financial interest(s): J.S., V.T., and M.P. are employees of BioVersys AG. M.H.M. is a member of the Scientific Advisory Board of Hexagon Bio and cofounder of Design Pharmaceuticals.
Figures




References
-
- Murray C. J. L.; Ikuta K. S.; Sharara F.; Swetschinski L.; Robles Aguilar G.; Gray A.; Han C.; Bisignano C.; Rao P.; Wool E.; Johnson S. C.; Browne A. J.; Chipeta M. G.; Fell F.; Hackett S.; Haines-Woodhouse G.; Kashef Hamadani B. H.; Kumaran E. A. P.; McManigal B.; Achalapong S.; Agarwal R.; Akech S.; Albertson S.; Amuasi J.; Andrews J.; Aravkin A.; Ashley E.; Babin F.-X.; Bailey F.; Baker S.; Basnyat B.; Bekker A.; Bender R.; Berkley J. A.; Bethou A.; Bielicki J.; Boonkasidecha S.; Bukosia J.; Carvalheiro C.; Castaneda-Orjuela C.; Chansamouth V.; Chaurasia S.; Chiurchiu S.; Chowdhury F.; Clotaire Donatien R.; Cook A. J.; Cooper B.; Cressey T. R.; Criollo-Mora E.; Cunningham M.; Darboe S.; Day N. P. J.; De Luca M.; Dokova K.; Dramowski A.; Dunachie S. J.; Duong Bich T.; Eckmanns T.; Eibach D.; Emami A.; Feasey N.; Fisher-Pearson N.; Forrest K.; Garcia C.; Garrett D.; Gastmeier P.; Giref A. Z.; Greer R. C.; Gupta V.; Haller S.; Haselbeck A.; Hay S. I.; Holm M.; Hopkins S.; Hsia Y.; Iregbu K. C.; Jacobs J.; Jarovsky D.; Javanmardi F.; Jenney A. W. J.; Khorana M.; Khusuwan S.; Kissoon N.; Kobeissi E.; Kostyanev T.; Krapp F.; Krumkamp R.; Kumar A.; Kyu H. H.; Lim C.; Lim K.; Limmathurotsakul D.; Loftus M. J.; Lunn M.; Ma J.; Manoharan A.; Marks F.; May J.; Mayxay M.; Mturi N.; Munera-Huertas T.; Musicha P.; Musila L. A.; Mussi-Pinhata M. M.; Naidu R. N.; Nakamura T.; Nanavati R.; Nangia S.; Newton P.; Ngoun C.; Novotney A.; Nwakanma D.; Obiero C. W.; Ochoa T. J.; Olivas-Martinez A.; Olliaro P.; Ooko E.; Ortiz-Brizuela E.; Ounchanum P.; Pak G. D.; Paredes J. L.; Peleg A. Y.; Perrone C.; Phe T.; Phommasone K.; Plakkal N.; Ponce-de-Leon A.; Raad M.; Ramdin T.; Rattanavong S.; Riddell A.; Roberts T.; Robotham J. V.; Roca A.; Rosenthal V. D.; Rudd K. E.; Russell N.; Sader H. S.; Saengchan W.; Schnall J.; Scott J. A. G.; Seekaew S.; Sharland M.; Shivamallappa M.; Sifuentes-Osornio J.; Simpson A. J.; Steenkeste N.; Stewardson A. J.; Stoeva T.; Tasak N.; Thaiprakong A.; Thwaites G.; Tigoi C.; Turner C.; Turner P.; van Doorn H. R.; Velaphi S.; Vongpradith A.; Vongsouvath M.; Vu H.; Walsh T.; Walson J. L.; Waner S.; Wangrangsimakul T.; Wannapinij P.; Wozniak T.; Young Sharma T. E. M. W.; Yu K. C.; Zheng P.; Sartorius B.; Lopez A. D.; Stergachis A.; Moore C.; Dolecek C.; Naghavi M. Global Burden of Bacterial Antimicrobial Resistance in 2019: A Systematic Analysis. Lancet 2022, 399 (10325), 629–655. 10.1016/S0140-6736(21)02724-0. - DOI - PMC - PubMed
-
- O’Neill J.Antimicrobial Resistance: Tackling a Crisis for the Health and Wealth of Nations; Review on Antimicrobial Resistance: London, 2014.
-
- O’Neill J.Tackling Drug-Resistant Infections Globally: Final Report and Recommendations; Review on Antimicrobial Resistance: London, 2016.
-
- Centers for Disease Control and Prevention; National Center for Emerging and Zoonotic Infectious Diseases; Division of Healthcare Quality Promotion . COVID-19: U.S. Impact on Antimicrobial Resistance, Special Report 2022; Centers for Disease Control and Prevention: Atlanta, GA, 2022.10.15620/cdc:117915. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

