Genomic landscape of Down syndrome-associated acute lymphoblastic leukemia
- PMID: 37001051
- PMCID: PMC10352600
- DOI: 10.1182/blood.2023019765
Genomic landscape of Down syndrome-associated acute lymphoblastic leukemia
Abstract
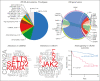
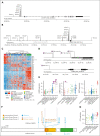
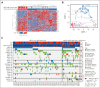
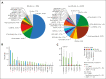
Trisomy 21, the genetic cause of Down syndrome (DS), is the most common congenital chromosomal anomaly. It is associated with a 20-fold increased risk of acute lymphoblastic leukemia (ALL) during childhood and results in distinctive leukemia biology. To comprehensively define the genomic landscape of DS-ALL, we performed whole-genome sequencing and whole-transcriptome sequencing (RNA-Seq) on 295 cases. Our integrated genomic analyses identified 15 molecular subtypes of DS-ALL, with marked enrichment of CRLF2-r, IGH::IGF2BP1, and C/EBP altered (C/EBPalt) subtypes compared with 2257 non-DS-ALL cases. We observed abnormal activation of the CEBPD, CEBPA, and CEBPE genes in 10.5% of DS-ALL cases via a variety of genomic mechanisms, including chromosomal rearrangements and noncoding mutations leading to enhancer hijacking. A total of 42.3% of C/EBP-activated DS-ALL also have concomitant FLT3 point mutations or insertions/deletions, compared with 4.1% in other subtypes. CEBPD overexpression enhanced the differentiation of mouse hematopoietic progenitor cells into pro-B cells in vitro, particularly in a DS genetic background. Notably, recombination-activating gene-mediated somatic genomic abnormalities were common in DS-ALL, accounting for a median of 27.5% of structural alterations, compared with 7.7% in non-DS-ALL. Unsupervised hierarchical clustering analyses of CRLF2-rearranged DS-ALL identified substantial heterogeneity within this group, with the BCR::ABL1-like subset linked to an inferior event-free survival, even after adjusting for known clinical risk factors. These results provide important insights into the biology of DS-ALL and point to opportunities for targeted therapy and treatment individualization.
© 2023 by The American Society of Hematology.
Conflict of interest statement
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Figures


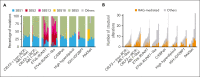
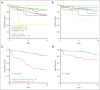
Comment in
-
All about Down syndrome ALL.Blood. 2023 Jul 13;142(2):126-128. doi: 10.1182/blood.2023020508. Blood. 2023. PMID: 37440267 No abstract available.
References
-
- Hasle H, Clemmensen IH, Mikkelsen M. Risks of leukaemia and solid tumours in individuals with Down’s syndrome. Lancet. 2000;355(9199):165–169. - PubMed
Publication types
MeSH terms
Grants and funding
- U24 CA196173/CA/NCI NIH HHS/United States
- S10 OD025240/OD/NIH HHS/United States
- R01 CA249867/CA/NCI NIH HHS/United States
- S10 OD023469/OD/NIH HHS/United States
- P30 EY002520/EY/NEI NIH HHS/United States
- U10 CA098543/CA/NCI NIH HHS/United States
- S10 RR024574/RR/NCRR NIH HHS/United States
- U10 CA180899/CA/NCI NIH HHS/United States
- U10 CA180886/CA/NCI NIH HHS/United States
- R01 GM134382/GM/NIGMS NIH HHS/United States
- U10 CA098413/CA/NCI NIH HHS/United States
- R03 HD103908/HD/NICHD NIH HHS/United States
- U24 CA114766/CA/NCI NIH HHS/United States
- P30 ES030285/ES/NIEHS NIH HHS/United States
- P30 CA125123/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

