Sulfur, sterol and trehalose metabolism in the deep-sea hydrocarbon seep tubeworm Lamellibrachia luymesi
- PMID: 37020304
- PMCID: PMC10077716
- DOI: 10.1186/s12864-023-09267-8
Sulfur, sterol and trehalose metabolism in the deep-sea hydrocarbon seep tubeworm Lamellibrachia luymesi
Abstract
Background: Lamellibrachia luymesi dominates cold sulfide-hydrocarbon seeps and is known for its ability to consume bacteria for energy. The symbiotic relationship between tubeworms and bacteria with particular adaptations to chemosynthetic environments has received attention. However, metabolic studies have primarily focused on the mechanisms and pathways of the bacterial symbionts, while studies on the animal hosts are limited.
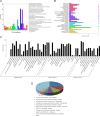
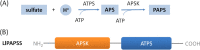
Results: Here, we sequenced the transcriptome of L. luymesi and generated a transcriptomic database containing 79,464 transcript sequences. Based on GO and KEGG annotations, we identified transcripts related to sulfur metabolism, sterol biosynthesis, trehalose synthesis, and hydrolysis. Our in-depth analysis identified sulfation pathways in L. luymesi, and sulfate activation might be an important detoxification pathway for promoting sulfur cycling, reducing byproducts of sulfide metabolism, and converting sulfur compounds to sulfur-containing organics, which are essential for symbiotic survival. Moreover, sulfide can serve directly as a sulfur source for cysteine synthesis in L. luymesi. The existence of two pathways for cysteine synthesis might ensure its participation in the formation of proteins, heavy metal detoxification, and the sulfide-binding function of haemoglobin. Furthermore, our data suggested that cold-seep tubeworm is capable of de novo sterol biosynthesis, as well as incorporation and transformation of cycloartenol and lanosterol into unconventional sterols, and the critical enzyme involved in this process might have properties similar to those in the enzymes from plants or fungi. Finally, trehalose synthesis in L. luymesi occurs via the trehalose-6-phosphate synthase (TPS) and trehalose-6-phosphate phosphatase (TPP) pathways. The TPP gene has not been identified, whereas the TPS gene encodes a protein harbouring conserved TPS/OtsA and TPP/OtsB domains. The presence of multiple trehalases that catalyse trehalose hydrolysis could indicate the different roles of trehalase in cold-seep tubeworms.
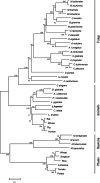
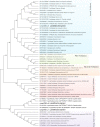
Conclusions: We elucidated several molecular pathways of sulfate activation, cysteine and cholesterol synthesis, and trehalose metabolism. Contrary to the previous analysis, two pathways for cysteine synthesis and the cycloartenol-C-24-methyltransferase gene were identified in animals for the first time. The present study provides new insights into particular adaptations to chemosynthetic environments in L. luymesi and can serve as the basis for future molecular studies on host-symbiont interactions and biological evolution.
Keywords: Adaptation to deep-sea chemosynthetic environment; Cholesterol; Cold seep; Cycloartenol-C-24-methyltransferase; Cysteine; Fungi; Sulfate; Transcriptome; Trehalase.
© 2023. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures





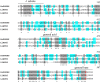
References
-
- Gardiner S, Hourdez S. On the occurrence of the vestimentiferan tube worm Lamellibrachia luymesi van de Land and Norrevang, 1975 (Annelida: Pogonophora) in hydrocarbon seep communities in the Gulf of Mexico. Biological Society of Washington. 2003;116:380–394.
-
- Flores JF, Fisher CR, Carney SL, Green BN, Freytag JK, Schaeffer SW, Royer WE., Jr Sulfide binding is mediated by zinc ions discovered in the crystal structure of a hydrothermal vent tubeworm hemoglobin. Proc Natl Acad Sci U S A. 2005;102(8):2713–2718. doi: 10.1073/pnas.0407455102. - DOI - PMC - PubMed
-
- Bailly X, Jollivet D, Vanin S, Deutsch J, Zal F, Lallier F, Toulmond A. Evolution of the sulfide-binding function within the globin multigenic family of the deep-sea hydrothermal vent tubeworm Riftia pachyptila. Mol Biol Evol. 2002;19(9):1421–1433. doi: 10.1093/oxfordjournals.molbev.a004205. - DOI - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

