Elements of divergence in germline determination in closely related species
- PMID: 37020963
- PMCID: PMC10068562
- DOI: 10.1016/j.isci.2023.106402
Elements of divergence in germline determination in closely related species
Abstract
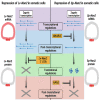
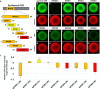
Evolutionary transitions are particularly important in development of the germ line, cells which directly impact sexual reproduction. Differences in the primordial germ cells (PGCs) of two sea urchin species were examined here by stage-matched, integrated, single cell RNA-seq (scRNA-seq) datasets. Even though both species rely on inherited mechanisms to specify their germ line, this analysis revealed a variety of differences in germline gene expression, including a broader expression of the germline factor Nanos2 (Nan2) in Lytechinus variegatus (Lv) compared to Strongylocentrotus purpuratus (Sp). In Sp, Nan2 mRNA expression is highly restricted to the PGCs by a lability element in its 3'UTR, which is lacking in the mRNA of Lv-Nan2, thus explaining the difference. We discovered that the Lv-Nan2 3'UTR instead leads to its specific translation in the PGCs. The results emphasize that regulatory mechanisms resulting in germline specification rely greatly on post-transcriptional restrictions of key gene products.
Keywords: Cell biology; Evolutionary biology.
© 2023 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






Similar articles
-
Functional annotation of a hugely expanded nanos repertoire in Lytechinus variegatus, the green sea urchin.Mol Reprod Dev. 2023 May;90(5):310-322. doi: 10.1002/mrd.23684. Epub 2023 Apr 11. Mol Reprod Dev. 2023. PMID: 37039283 Free PMC article.
-
The 3'UTR of nanos2 directs enrichment in the germ cell lineage of the sea urchin.Dev Biol. 2013 May 1;377(1):275-83. doi: 10.1016/j.ydbio.2013.01.019. Epub 2013 Jan 25. Dev Biol. 2013. PMID: 23357540 Free PMC article.
-
Transient translational quiescence in primordial germ cells.Development. 2017 Apr 1;144(7):1201-1210. doi: 10.1242/dev.144170. Epub 2017 Feb 24. Development. 2017. PMID: 28235822 Free PMC article.
-
Lessons for inductive germline determination.Mol Reprod Dev. 2013 Aug;80(8):590-609. doi: 10.1002/mrd.22151. Epub 2013 Feb 28. Mol Reprod Dev. 2013. PMID: 23450642 Free PMC article. Review.
-
The germline: familiar and newly uncovered properties.Annu Rev Genet. 1994;28:309-24. doi: 10.1146/annurev.ge.28.120194.001521. Annu Rev Genet. 1994. PMID: 7893129 Review.
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

