Genomic adaptation to extreme climate conditions in beef cattle as a consequence of cross-breeding program
- PMID: 37024818
- PMCID: PMC10080750
- DOI: 10.1186/s12864-023-09235-2
Genomic adaptation to extreme climate conditions in beef cattle as a consequence of cross-breeding program
Erratum in
-
Correction: Genomic adaptation to extreme climate conditions in beef cattle as a consequence of cross-breeding program.BMC Genomics. 2024 Nov 20;25(1):1117. doi: 10.1186/s12864-024-11042-2. BMC Genomics. 2024. PMID: 39567918 Free PMC article. No abstract available.
Abstract
Background: Understanding the evolutionary forces related to climate changes that have been shaped genetic variation within species has long been a fundamental pursuit in biology. In this study, we generated whole-genome sequence (WGS) data from 65 cross-bred and 45 Mongolian cattle. Together with 62 whole-genome sequences from world-wide cattle populations, we estimated the genetic diversity and population genetic structure of cattle populations. In addition, we performed comparative population genomics analyses to explore the genetic basis underlying variation in the adaptation to cold climate and immune response in cross-bred cattle located in the cold region of China. To elucidate genomic signatures that underlie adaptation to cold climate, we performed three statistical measurements, fixation index (FST), log2 nucleotide diversity (θπ ratio) and cross population composite likelihood ratio (XP-CLR), and further investigated the results to identify genomic regions under selection for cold adaptation and immune response-related traits.
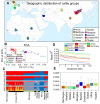
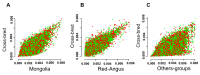
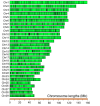
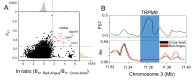
Results: By generating WGS data, we investigated the population genetic structure and phylogenetic relationship of studied cattle populations. The results revealed clustering of cattle groups in agreement with their geographic distribution. We detected noticeable genetic diversity between indigenous cattle ecotypes and commercial populations. Analysis of population structure demonstrated evidence of shared genetic ancestry between studied cross-bred population and both Red-Angus and Mongolian breeds. Among all studied cattle populations, the highest and lowest levels of linkage disequilibrium (LD) per Kb were detected in Holstein and Rashoki populations (ranged from ~ 0.54 to 0.73, respectively). Our search for potential genomic regions under selection in cross-bred cattle revealed several candidate genes related with immune response and cold shock protein on multiple chromosomes. We identified some adaptive introgression genes with greater than expected contributions from Mongolian ancestry into Molgolian x Red Angus composites such as TRPM8, NMUR1, PRKAA2, SMTNL2 and OXR1 that are involved in energy metabolism and metabolic homeostasis. In addition, we detected some candidate genes probably associated with immune response-related traits.
Conclusion: The study identified candidate genes involved in responses to cold adaptation and immune response in cross-bred cattle, including new genes or gene pathways putatively involved in these adaptations. The identification of these genes may clarify the molecular basis underlying adaptation to extreme environmental climate and as such they might be used in cattle breeding programs to select more efficient breeds for cold climate regions.
Keywords: Cattle; Cold adaptation; TRPM8; Whole-genome sequence.
© 2023. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
Similar articles
-
Uncovering genomic diversity and signatures of selection in red Angus × Chinese red steppe crossbred cattle population.Sci Rep. 2025 Apr 15;15(1):12977. doi: 10.1038/s41598-025-98346-9. Sci Rep. 2025. PMID: 40234714 Free PMC article.
-
Genetic diversity and signatures of selection for heat tolerance and immune response in Iranian native chickens.BMC Genomics. 2022 Mar 22;23(1):224. doi: 10.1186/s12864-022-08434-7. BMC Genomics. 2022. PMID: 35317755 Free PMC article.
-
Assessing genomic diversity and signatures of selection in Jiaxian Red cattle using whole-genome sequencing data.BMC Genomics. 2021 Jan 9;22(1):43. doi: 10.1186/s12864-020-07340-0. BMC Genomics. 2021. PMID: 33421990 Free PMC article. Review.
-
Genome-wide scan for selection signatures reveals novel insights into the adaptive capacity characteristics in three Chinese cattle breeds.BMC Genomics. 2025 Feb 28;26(1):206. doi: 10.1186/s12864-025-11328-z. BMC Genomics. 2025. PMID: 40021973 Free PMC article.
-
Genome-wide selection signatures detection in Shanghai Holstein cattle population identified genes related to adaption, health and reproduction traits.BMC Genomics. 2021 Oct 15;22(1):747. doi: 10.1186/s12864-021-08042-x. BMC Genomics. 2021. PMID: 34654366 Free PMC article. Review.
Cited by
-
Whole-Genome Selective Scans Detect Genes Associated with Cashmere Traits and Climatic Adaptation in Cashmere Goats (Capra hircus) in China.Genes (Basel). 2025 Feb 27;16(3):292. doi: 10.3390/genes16030292. Genes (Basel). 2025. PMID: 40149444 Free PMC article.
-
Uncovering genomic diversity and signatures of selection in red Angus × Chinese red steppe crossbred cattle population.Sci Rep. 2025 Apr 15;15(1):12977. doi: 10.1038/s41598-025-98346-9. Sci Rep. 2025. PMID: 40234714 Free PMC article.
-
Functional analysis of microorganisms and metabolites in the cecum of different sheep populations and their effects on production traits.Front Microbiol. 2024 Sep 16;15:1437250. doi: 10.3389/fmicb.2024.1437250. eCollection 2024. Front Microbiol. 2024. PMID: 39351299 Free PMC article.
-
scalepopgen: Bioinformatic Workflow Resources Implemented in Nextflow for Comprehensive Population Genomic Analyses.Mol Biol Evol. 2024 Apr 2;41(4):msae057. doi: 10.1093/molbev/msae057. Mol Biol Evol. 2024. PMID: 38507648 Free PMC article.
-
Potential Genetic Markers Associated with Environmental Adaptability in Herbivorous Livestock.Animals (Basel). 2025 Mar 5;15(5):748. doi: 10.3390/ani15050748. Animals (Basel). 2025. PMID: 40076029 Free PMC article. Review.
References
-
- Hume DA, Whitelaw CBA, Archibald AL. The future of animal production: improving productivity and sustainability. J Agric Sci. 2011;149:9–16.
-
- Makkar HPS. Review: Feed demand landscape and implications of food-not feed strategy for food security and climate change, Anim.12 (2018) 1744–1754. - PubMed
-
- FAO., 2020. Food Outlook – Biannual Report on Global Food Markets. Food and Agriculture Organization of the United Nation, Rome, Italy. https://www.fao.org/3/ca9509en/ca9509en.pdf.
-
- Greenwood PL. Review: an overview of beef production from pasture and feedlot globally, as demand for beef and the need for sustainable practices increase. Anim. 2021;15:100295. - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

