Molecular subtypes predict therapeutic responses and identifying and validating diagnostic signatures based on machine learning in chronic myeloid leukemia
- PMID: 37024911
- PMCID: PMC10080819
- DOI: 10.1186/s12935-023-02905-x
Molecular subtypes predict therapeutic responses and identifying and validating diagnostic signatures based on machine learning in chronic myeloid leukemia
Abstract
Chronic myeloid leukemia (CML) is a hematological tumor derived from hematopoietic stem cells. The aim of this study is to analyze the biological characteristics and identify the diagnostic markers of CML. We obtained the expression profiles from the Gene Expression Omnibus (GEO) database and identified 210 differentially expressed genes (DEGs) between CML and normal samples. These DEGs are mainly enriched in immune-related pathways such as Th1 and Th2 cell differentiation, primary immunodeficiency, T cell receptor signaling pathway, antigen processing and presentation pathways. Based on these DEGs, we identified two molecular subtypes using a consensus clustering algorithm. Cluster A was an immunosuppressive phenotype with reduced immune cell infiltration and significant activation of metabolism-related pathways such as reactive oxygen species, glycolysis and mTORC1; Cluster B was an immune activating phenotype with increased infiltration of CD4 + and CD8 + T cells and NK cells, and increased activation of signaling pathways such as interferon gamma (IFN-γ) response, IL6-JAK-STAT3 and inflammatory response. Drug prediction results showed that patients in Cluster B had a higher therapeutic response to anti-PD-1 and anti-CTLA4 and were more sensitive to imatinib, nilotinib and dasatinib. Support Vector Machine Recursive Feature Elimination (SVM-RFE), Least Absolute Shrinkage Selection Operator (LASSO) and Random Forest (RF) algorithms identified 4 CML diagnostic genes (HDC, SMPDL3A, IRF4 and AQP3), and the risk score model constructed by these genes improved the diagnostic accuracy. We further validated the diagnostic value of the 4 genes and the risk score model in a clinical cohort, and the risk score can be used in the differential diagnosis of CML and other hematological malignancies. The risk score can also be used to identify molecular subtypes and predict response to imatinib treatment. These results reveal the characteristics of immunosuppression and metabolic reprogramming in CML patients, and the identification of molecular subtypes and biomarkers provides new ideas and insights for the clinical diagnosis and treatment.
Keywords: Chronic myeloid leukemia; Diagnosis; Machine learning; Therapeutic response; Tumor microenvironment.
© 2023. The Author(s).
Conflict of interest statement
The authors have no competing interests to disclose.
Figures




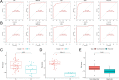



Similar articles
-
The therapeutic and biomarker significance of ferroptosis in chronic myeloid leukemia.Front Immunol. 2024 Jul 4;15:1402669. doi: 10.3389/fimmu.2024.1402669. eCollection 2024. Front Immunol. 2024. PMID: 39026664 Free PMC article.
-
Identification and validation of hub genes and molecular classifications associated with chronic myeloid leukemia.Front Immunol. 2024 Jan 12;14:1297886. doi: 10.3389/fimmu.2023.1297886. eCollection 2023. Front Immunol. 2024. PMID: 38283355 Free PMC article.
-
Autophagy crosstalk with the immune microenvironment in chronic myeloid leukemia and serves as a biomarker for diagnosis and progression.Front Immunol. 2025 May 28;16:1570903. doi: 10.3389/fimmu.2025.1570903. eCollection 2025. Front Immunol. 2025. PMID: 40503228 Free PMC article.
-
Identification of hub genes and their correlation with immune infiltration in coronary artery disease through bioinformatics and machine learning methods.J Thorac Dis. 2022 Jul;14(7):2621-2634. doi: 10.21037/jtd-22-632. J Thorac Dis. 2022. PMID: 35928610 Free PMC article.
-
Identification of diagnostic signatures associated with immune infiltration in Alzheimer's disease by integrating bioinformatic analysis and machine-learning strategies.Front Aging Neurosci. 2022 Jul 29;14:919614. doi: 10.3389/fnagi.2022.919614. eCollection 2022. Front Aging Neurosci. 2022. PMID: 35966794 Free PMC article.
Cited by
-
Profiling of miRNAs and their interfering targets in peripheral blood mononuclear cells from patients with chronic myeloid leukaemia.Front Oncol. 2023 Jul 5;13:1173970. doi: 10.3389/fonc.2023.1173970. eCollection 2023. Front Oncol. 2023. PMID: 37476380 Free PMC article.
-
Machine learning-based identification of efficient and restrictive physiological subphenotypes in acute respiratory distress syndrome.Intensive Care Med Exp. 2025 Mar 1;13(1):29. doi: 10.1186/s40635-025-00737-9. Intensive Care Med Exp. 2025. PMID: 40024962 Free PMC article.
-
Immune checkpoints PD1/PDL1, TIM3/GAL9 and key immune mediators landscape reveal differential expression dynamics on imatinib response in chronic myeloid leukemia.Ann Hematol. 2024 Dec;103(12):5249-5260. doi: 10.1007/s00277-024-06074-3. Epub 2024 Nov 7. Ann Hematol. 2024. PMID: 39505795
-
Global research trends on biomarkers for cancer immunotherapy: Visualization and bibliometric analysis.Hum Vaccin Immunother. 2025 Dec;21(1):2435598. doi: 10.1080/21645515.2024.2435598. Epub 2025 Jan 8. Hum Vaccin Immunother. 2025. PMID: 39773010 Free PMC article.
-
Application of artificial intelligence in chronic myeloid leukemia (CML) disease prediction and management: a scoping review.BMC Cancer. 2024 Aug 20;24(1):1026. doi: 10.1186/s12885-024-12764-y. BMC Cancer. 2024. PMID: 39164653 Free PMC article.
References
-
- Iqbal Z, et al. Sensitive detection of pre-existing BCR-ABL kinase domain mutations in CD34+ cells of newly diagnosed chronic-phase chronic myeloid leukemia patients is associated with imatinib resistance: implications in the post-imatinib era. PLoS ONE. 2013;8:e55717. doi: 10.1371/journal.pone.0055717. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

