Chromosome-level, nanopore-only genome and allele-specific DNA methylation of Pallas's cat, Otocolobus manul
- PMID: 37025970
- PMCID: PMC10071556
- DOI: 10.1093/nargab/lqad033
Chromosome-level, nanopore-only genome and allele-specific DNA methylation of Pallas's cat, Otocolobus manul
Abstract
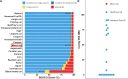
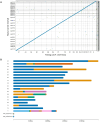
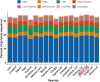
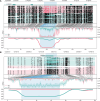
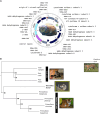
Pallas's cat, or the manul cat (Otocolobus manul), is a small felid native to the grasslands and steppes of central Asia. Population strongholds in Mongolia and China face growing challenges from climate change, habitat fragmentation, poaching, and other sources. These threats, combined with O. manul's zoo collection popularity and value in evolutionary biology, necessitate improvement of species genomic resources. We used standalone nanopore sequencing to assemble a 2.5 Gb, 61-contig nuclear assembly and 17097 bp mitogenome for O. manul. The primary nuclear assembly had 56× sequencing coverage, a contig N50 of 118 Mb, and a 94.7% BUSCO completeness score for Carnivora-specific genes. High genome collinearity within Felidae permitted alignment-based scaffolding onto the fishing cat (Prionailurus viverrinus) reference genome. Manul contigs spanned all 19 felid chromosomes with an inferred total gap length of less than 400 kilobases. Modified basecalling and variant phasing produced an alternate pseudohaplotype assembly and allele-specific DNA methylation calls; 61 differentially methylated regions were identified between haplotypes. Nearest features included classical imprinted genes, non-coding RNAs, and putative novel imprinted loci. The assembled mitogenome successfully resolved existing discordance between Felinae nuclear and mtDNA phylogenies. All assembly drafts were generated from 158 Gb of sequence using seven minION flow cells.
© The Author(s) 2023. Published by Oxford University Press on behalf of NAR Genomics and Bioinformatics.
Figures


Similar articles
-
Four new genome sequences of the Pallas's cat (Otocolobus manul): an insight into the patterns of within-species variability.Front Genet. 2024 Dec 9;15:1463774. doi: 10.3389/fgene.2024.1463774. eCollection 2024. Front Genet. 2024. PMID: 39720181 Free PMC article.
-
An intraerythrocytic small piroplasm in wild-caught Pallas's cats (Otocolobus manul) from Mongolia.J Wildl Dis. 2003 Apr;39(2):424-30. doi: 10.7589/0090-3558-39.2.424. J Wildl Dis. 2003. PMID: 12910772
-
Ecological niche models reveal divergent habitat use of Pallas's cat in the Eurasian cold steppes.Ecol Evol. 2022 Dec 14;12(12):e9624. doi: 10.1002/ece3.9624. eCollection 2022 Dec. Ecol Evol. 2022. PMID: 36532134 Free PMC article.
-
Dietary Differentiation Mitigates Interspecific Interference Competition Between Sympatric Pallas's Cats (Otocolobus manul) and Red Foxes (Vulpes vulpes).Animals (Basel). 2025 Apr 29;15(9):1267. doi: 10.3390/ani15091267. Animals (Basel). 2025. PMID: 40362082 Free PMC article.
-
The genome of Przewalski's horse (Equus ferus przewalskii).bioRxiv [Preprint]. 2024 Feb 28:2024.02.20.581252. doi: 10.1101/2024.02.20.581252. bioRxiv. 2024. Update in: G3 (Bethesda). 2024 Aug 7;14(8):jkae113. doi: 10.1093/g3journal/jkae113. PMID: 38464182 Free PMC article. Updated. Preprint.
Cited by
-
A high-quality Oxford Nanopore assembly of the hourglass dolphin (Lagenorhynchus cruciger) genome.G3 (Bethesda). 2025 May 8;15(5):jkaf044. doi: 10.1093/g3journal/jkaf044. G3 (Bethesda). 2025. PMID: 40036857 Free PMC article.
-
Four new genome sequences of the Pallas's cat (Otocolobus manul): an insight into the patterns of within-species variability.Front Genet. 2024 Dec 9;15:1463774. doi: 10.3389/fgene.2024.1463774. eCollection 2024. Front Genet. 2024. PMID: 39720181 Free PMC article.
-
The Genome of the Soybean Gall Midge ( Resseliella maxima ).bioRxiv [Preprint]. 2023 Feb 12:2023.02.10.528044. doi: 10.1101/2023.02.10.528044. bioRxiv. 2023. Update in: G3 (Bethesda). 2023 Apr 11;13(4):jkad046. doi: 10.1093/g3journal/jkad046. PMID: 36798210 Free PMC article. Updated. Preprint.
-
Statistical framework for calling allelic imbalance in high-throughput sequencing data.Nat Commun. 2025 Feb 18;16(1):1739. doi: 10.1038/s41467-024-55513-2. Nat Commun. 2025. PMID: 39966391 Free PMC article.
-
The genome of the soybean gall midge (Resseliella maxima).G3 (Bethesda). 2023 Apr 11;13(4):jkad046. doi: 10.1093/g3journal/jkad046. G3 (Bethesda). 2023. PMID: 36861345 Free PMC article.
References
-
- Ross S., Barashkova A., Dhendup T., Munkhtsog B., Smelansky I., Barclay D., moqanaki E. Otocolobus Manul. 2019; 10 February 2023, date last accessedhttp://www.10.2305/iucn.uk.2020-2.rlts.t15640a180145377.en.
-
- Gittleman J.L. Heptner, V.G. and Sludskii, A.A. 1992. Mammals of the soviet union. volume II, part 2. Carnivora (hyaenas and cats). Smithsonian Institution Libraries and National Science Foundation. J. Mammal. 1993; 74:510–511.
-
- Murdoch J., Tserendorj M., Reading R. Pallas’ cat ecology and conservation in the semi-desert steppes of mongolia. CAT News. 2006; 45:18–19.
-
- BBC The grumpiest cat in the world. 2022; 10 February 2023, date last accessedhttps://www.bbc.co.uk/programmes/p0cz5p0y.
-
- Ross S., Munkhtsog B., Harris S. Dietary composition, plasticity, and prey selection of Pallas's cats. J. Mammal. 2010; 91:811–817.
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

