Nanopore sequencing reveals methylation changes associated with obesity in circulating cell-free DNA from Göttingen Minipigs
- PMID: 37032646
- PMCID: PMC10088973
- DOI: 10.1080/15592294.2023.2199374
Nanopore sequencing reveals methylation changes associated with obesity in circulating cell-free DNA from Göttingen Minipigs
Abstract
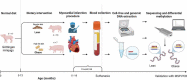
Profiling of circulating cell-free DNA (cfDNA) by tissue-specific base modifications, such as 5-methylcytosines (5mC), may enable the monitoring of ongoing pathophysiological processes. Nanopore sequencing allows genome-wide 5mC detection in cfDNA without bisulphite conversion. The aims of this study were: i) to find differentially methylated regions (DMRs) of cfDNA associated with obesity in Göttingen minipigs using Nanopore sequencing, ii) to validate a subset of the DMRs using methylation-specific PCR (MSP-PCR), and iii) to compare the cfDNA DMRs with those from whole blood genomic DNA (gDNA). Serum cfDNA and gDNA were obtained from 10 lean and 7 obese Göttingen Minipigs both with experimentally induced myocardial infarction and sequenced using Oxford Nanopore MinION. A total of 1,236 cfDNA DMRs (FDR<0.01) were associated with obesity. In silico analysis showed enrichment of the adipocytokine signalling, glucagon signalling, and cellular glucose homoeostasis pathways. A strong cfDNA DMR was discovered in PPARGC1B, a gene linked to obesity and type 2 diabetes. The DMR was validated using MSP-PCR and correlated significantly with body weight (P < 0.05). No DMRs intersected between cfDNA and gDNA, suggesting that cfDNA originates from body-wide shedding of DNA. In conclusion, nanopore sequencing detected differential methylation in minute quantities (0.1-1 ng/µl) of cfDNA. Future work should focus on translation into human and comparing 5mC from somatic tissues to pinpoint the exact location of pathology.
Keywords: Cell-free DNA; Nanopore sequencing; diagnostics; epigenetics; methylation; obesity.
Plain language summary
Oxford nanopore sequencing can reveal changes in methylation patterns associated with obesity in minute quantities of cell-free DNA from serum.Bisulphite conversion and methylation-specific PCR can be used to validate differentially methylated regions in cell-free DNA.A differentially methylated region in an intronic region of the PPARGC1B gene was found associated with obesity.Differentially methylated regions in cell-free DNA could be useful as early risk markers of certain diseases and pathologies.
Conflict of interest statement
No potential conflict of interest was reported by the authors.
Figures





Similar articles
-
Genome-wide methylation profiling of maternal cell-free DNA using methylated DNA sequencing (MeD-seq) indicates a placental and immune-cell signature.Eur J Clin Invest. 2025 Mar;55(3):e14363. doi: 10.1111/eci.14363. Epub 2024 Nov 26. Eur J Clin Invest. 2025. PMID: 39589853 Free PMC article.
-
Discovery and Validation of Methylation Signatures in Circulating Cell-Free DNA for the Detection of Colorectal Cancer.Biomolecules. 2024 Aug 13;14(8):996. doi: 10.3390/biom14080996. Biomolecules. 2024. PMID: 39199384 Free PMC article.
-
Distinct methylome profile of cfDNA in AMI patients reveals significant alteration in cAMP signaling pathway genes regulating cardiac muscle contraction.Clin Epigenetics. 2024 Oct 16;16(1):144. doi: 10.1186/s13148-024-01755-2. Clin Epigenetics. 2024. PMID: 39415189 Free PMC article.
-
Computational challenges in detection of cancer using cell-free DNA methylation.Comput Struct Biotechnol J. 2021 Dec 7;20:26-39. doi: 10.1016/j.csbj.2021.12.001. eCollection 2022. Comput Struct Biotechnol J. 2021. PMID: 34976309 Free PMC article. Review.
-
DNA methylation biomarkers for noninvasive detection of triple-negative breast cancer using liquid biopsy.Int J Cancer. 2023 Mar 1;152(5):1025-1035. doi: 10.1002/ijc.34337. Epub 2022 Nov 8. Int J Cancer. 2023. PMID: 36305646 Review.
Cited by
-
Current Updates on the Understanding of the Role of DNA Methylation on Obesity.Diabetes Metab Syndr Obes. 2024 Aug 27;17:3177-3186. doi: 10.2147/DMSO.S471348. eCollection 2024. Diabetes Metab Syndr Obes. 2024. PMID: 39220797 Free PMC article. Review.
-
Impact of obesity on infarct size, circulating biomarkers, mitochondrial function and mortality in a Göttingen minipig myocardial infarct model.Lab Anim (NY). 2025 Apr;54(4):103-111. doi: 10.1038/s41684-025-01533-4. Epub 2025 Mar 27. Lab Anim (NY). 2025. PMID: 40164843 Free PMC article.
-
Mechanisms and technologies in cancer epigenetics.Front Oncol. 2025 Jan 7;14:1513654. doi: 10.3389/fonc.2024.1513654. eCollection 2024. Front Oncol. 2025. PMID: 39839798 Free PMC article. Review.
-
Circulating Cell-Free DNA in Metabolic Diseases.J Endocr Soc. 2025 Jan 15;9(2):bvaf006. doi: 10.1210/jendso/bvaf006. eCollection 2025 Jan 6. J Endocr Soc. 2025. PMID: 39850787 Free PMC article. Review.
References
-
- Peng X, Li H-D, Wu F-X, et al. Identifying the tissues-of-origin of circulating cell-free DNAs is a promising way in noninvasive diagnostics. Brief Bioinform. 2020;22(3):bbaa060. - PubMed
-
- Jylhävä J, Lehtimäki T, Jula A, et al. Circulating cell-free DNA is associated with cardiometabolic risk factors: the health 2000 survey. Atherosclerosis. 2014;233:268–271. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous
