Evolutionary and in silico guided development of novel peptide analogues for antibacterial activity against ESKAPE pathogens
- PMID: 37032813
- PMCID: PMC10073642
- DOI: 10.1016/j.crmicr.2023.100183
Evolutionary and in silico guided development of novel peptide analogues for antibacterial activity against ESKAPE pathogens
Abstract
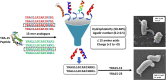
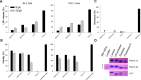
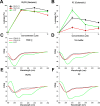
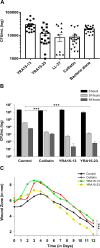
According to WHO, to combat the resistant strains, new effective anti-microbial agents are needed on an urgent basis and global researchers should focus their efforts and discovery programs on developing them against antibiotic-resistant pathogens or priority pathogens like ESKAPE. In this context, Cationic antimicrobial peptides (AMPs) are being explored extensively as promising next-generation antimicrobials due to their broad range, fast kinetics and multifunctional role. Despite recent advances, it is still a daunting challenge to identify and design a potent AMP with no cytotoxicity, but with broad specific antimicrobial activity, stability and efficacy under in vivo conditions in a cost-effective and robust manner. In this work, as a proof of concept, we designed novel potent AMPs using artificial intelligence based in silico programs. Shortlisted peptide sequences were synthesized using the fmoc chemistry approach, assessed their antimicrobial activity, cell selectivity, mode of action and in vivo efficacy using a series of experiments. The synthesized peptide analogues demonstrated their antimicrobial activity (MIC in the range of 2.5-80 μM) against bacteria. The identified potential lead molecules showed antibacterial activity in physiological conditions with no signs of cytotoxicity. We further tested the antimicrobial activity of peptide analogues for treating wounds infected with Pseudomonas aeruginosa in the mice burn wound model. In drug-development programs, the identification of lead antimicrobial agents is always challenging and involves screening a large number of molecules which is time-consuming and expensive. This work demonstrates the utility of artificial intelligence based in silico analysis programs in discovering novel antimicrobial agents in an economical, robust way.
Keywords: Antimicrobial peptides; Burn wound model; ESKAPE pathogens; Hydrophobicity and charge relationships; In silico designing; Peptide analoguess.
© 2023 The Author(s).
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper. The authors declare the following financial interests/personal relationships which may be considered as potential competing interests: Mukesh Pasupuleti reports a relationship with Council for Scientific and Industrial Research that includes: employment.
Figures

Similar articles
-
Rational Framework for the Design of Trp- and Arg-Rich Peptide Antibiotics Against Multidrug-Resistant Bacteria.Front Microbiol. 2022 May 23;13:889791. doi: 10.3389/fmicb.2022.889791. eCollection 2022. Front Microbiol. 2022. PMID: 35694289 Free PMC article.
-
Trematocine-derived antimicrobial peptides from the Antarctic fish Trematomus bernacchaii: potent antibacterial agents against ESKAPE pathogens.Front Microbiol. 2024 Aug 7;15:1447301. doi: 10.3389/fmicb.2024.1447301. eCollection 2024. Front Microbiol. 2024. PMID: 39171261 Free PMC article.
-
Cationicity Enhancement on the Hydrophilic Face of Ctriporin Significantly Reduces Its Hemolytic Activity and Improves the Antimicrobial Activity against Antibiotic-Resistant ESKAPE Pathogens.Toxins (Basel). 2024 Mar 18;16(3):156. doi: 10.3390/toxins16030156. Toxins (Basel). 2024. PMID: 38535822 Free PMC article.
-
The Resilience of Pseudomonas aeruginosa to Antibiotics and the Designing of Antimicrobial Peptides to Overcome Microbial Resistance.Curr Med Chem. 2022;30(1):72-103. doi: 10.2174/0929867329666220907100505. Curr Med Chem. 2022. PMID: 36082872 Review.
-
Short Peptides and Their Mimetics as Potent Antibacterial Agents and Antibiotic Adjuvants.ACS Chem Biol. 2021 Dec 17;16(12):2731-2745. doi: 10.1021/acschembio.1c00626. Epub 2021 Nov 15. ACS Chem Biol. 2021. PMID: 34779605 Review.
Cited by
-
Co-translational protein aggregation and ribosome stalling as a broad-spectrum antibacterial mechanism.Nat Commun. 2025 Feb 12;16(1):1561. doi: 10.1038/s41467-025-56873-z. Nat Commun. 2025. PMID: 39939597 Free PMC article.
-
Synthesis and structure-activity study of the antimicrobial lipopeptide brevibacillin.RSC Med Chem. 2024 Sep 25;15(12):4168-79. doi: 10.1039/d4md00612g. Online ahead of print. RSC Med Chem. 2024. PMID: 39371433 Free PMC article.
-
Potential application of Healitide-GP1, a novel antibacterial peptide, in wound healing: in vitro studies.Sci Rep. 2025 Aug 24;15(1):31078. doi: 10.1038/s41598-025-17002-4. Sci Rep. 2025. PMID: 40849541 Free PMC article.
-
Synthesis, characterization, quantum chemical modelling, molecular docking, in silico and in vitro assessment of 3-(2-bromo-5-fluorophenyl))-1-(thiophen-2-yl)prop-2-en-1-one.Sci Rep. 2024 Dec 1;14(1):29838. doi: 10.1038/s41598-024-79747-8. Sci Rep. 2024. PMID: 39617796 Free PMC article.
References
-
- de Oliveira Costa B, Franco O.L. Cryptic host defense peptides: multifaceted activity and prospects for medicinal chemistry. Curr. Top Med. Chem. 2020;20:1274–1290. - PubMed
LinkOut - more resources
Full Text Sources

