Identification of berberine as a potential therapeutic strategy for kidney clear cell carcinoma and COVID-19 based on analysis of large-scale datasets
- PMID: 37033923
- PMCID: PMC10076552
- DOI: 10.3389/fimmu.2023.1038651
Identification of berberine as a potential therapeutic strategy for kidney clear cell carcinoma and COVID-19 based on analysis of large-scale datasets
Abstract
Background: Regarding the global coronavirus disease 2019 (COVID)-19 pandemic, kidney clear cell carcinoma (KIRC) has acquired a higher infection probability and may induce fatal complications and death following COVID-19 infection. However, effective treatment strategies remain unavailable. Berberine exhibits significant antiviral and antitumour effects. Thus, this study aimed to provide a promising and reliable therapeutic strategy for clinical decision-making by exploring the therapeutic mechanism of berberine against KIRC/COVID-19.
Methods: Based on large-scale data analysis, the target genes, clinical risk, and immune and pharmacological mechanisms of berberine against KIRC/COVID-19 were systematically investigated.
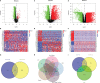
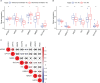
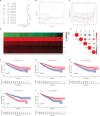
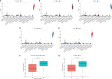
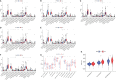
Results: In total, 1,038 and 12,992 differentially expressed genes (DEGs) of COVID-19 and KIRC, respectively, were verified from Gene Expression Omnibus and The Cancer Genome Atlas databases, respectively, and 489 berberine target genes were obtained from official websites. After intersecting, 26 genes were considered potential berberine therapeutic targets for KIRC/COVID-19. Berberine mechanism of action against KIRC/COVID-19 was revealed by protein-protein interaction, gene ontology, and Kyoto Encyclopedia of Genes and Genomes with terms including protein interaction, cell proliferation, viral carcinogenesis, and the PI3K/Akt signalling pathway. In COVID-19 patients, ACOX1, LRRK2, MMP8, SLC1A3, CPT1A, H2AC11, H4C8, and SLC1A3 were closely related to disease severity, and the general survival of KIRC patients was closely related to ACOX1, APP, CPT1A, PLK1, and TYMS. Additionally, the risk signature accurately and sensitively depicted the overall survival and patient survival status for KIRC. Numerous neutrophils were enriched in the immune system of COVID-19 patients, and the lives of KIRC patients were endangered due to significant immune cell infiltration. Molecular docking studies indicated that berberine binds strongly to target proteins.
Conclusion: This study demonstrated berberine as a potential treatment option in pharmacological, immunological, and clinical practice. Moreover, its therapeutic effects may provide potential and reliable treatment options for patients with KIRC/COVID-19.
Keywords: berberine; coronavirus disease 2019; immune mechanism; kidney clear cell carcinoma; molecular docking.
Copyright © 2023 Zheng, Li, Nie, Wang, Liang, Yang, Zheng and Zheng.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




Similar articles
-
Therapeutic Effects of Berberine against Urological Cancers: Biological Potentials Based on Cellular Mechanisms.Curr Mol Med. 2024;24(10):1282-1290. doi: 10.2174/0115665240263630231009050436. Curr Mol Med. 2024. PMID: 37933211 Review.
-
Identification of small molecule drugs and development of a novel autophagy-related prognostic signature for kidney renal clear cell carcinoma.Cancer Med. 2020 Oct;9(19):7034-7051. doi: 10.1002/cam4.3367. Epub 2020 Aug 11. Cancer Med. 2020. PMID: 32780567 Free PMC article.
-
BUB1-deficiency suppresses kidney renal clear cell carcinoma progression via the PI3K/Akt pathway: A bioinformatics-oriented validating study.Mol Cell Probes. 2025 Jun;81:102024. doi: 10.1016/j.mcp.2025.102024. Epub 2025 Mar 11. Mol Cell Probes. 2025. PMID: 40081509
-
SHMT as a Potential Therapeutic Target for Renal Cell Carcinoma.Front Biosci (Landmark Ed). 2023 Sep 12;28(9):196. doi: 10.31083/j.fbl2809196. Front Biosci (Landmark Ed). 2023. PMID: 37796681
-
Upregulated PPP1R14B is connected to cancer progression and immune infiltration in kidney renal clear cell carcinoma.Clin Transl Oncol. 2024 Jan;26(1):119-135. doi: 10.1007/s12094-023-03228-z. Epub 2023 Jun 1. Clin Transl Oncol. 2024. PMID: 37261660
Cited by
-
Recent Applications of Protoberberines as Privileged Starting Materials for the Development of Novel Broad-Spectrum Antiviral Agents: A Concise Review (2017-2023).ACS Pharmacol Transl Sci. 2023 Dec 28;7(1):48-71. doi: 10.1021/acsptsci.3c00292. eCollection 2024 Jan 12. ACS Pharmacol Transl Sci. 2023. PMID: 38230282 Free PMC article. Review.
-
Therapeutic Effects of Berberine against Urological Cancers: Biological Potentials Based on Cellular Mechanisms.Curr Mol Med. 2024;24(10):1282-1290. doi: 10.2174/0115665240263630231009050436. Curr Mol Med. 2024. PMID: 37933211 Review.
-
Unveiling the oncogenic significance of thymidylate synthase in human cancers.Am J Transl Res. 2024 Oct 15;16(10):5228-5247. doi: 10.62347/IRUZ1011. eCollection 2024. Am J Transl Res. 2024. PMID: 39544745 Free PMC article.
-
Bioinformatics and system biology approach to discover the common pathogenetic processes between COVID-19 and chronic hepatitis B.PLoS One. 2025 May 23;20(5):e0323708. doi: 10.1371/journal.pone.0323708. eCollection 2025. PLoS One. 2025. PMID: 40408617 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

