PTPMT1 regulates mitochondrial death through the SLC25A6-NDUFS2 axis in pancreatic cancer cells
- PMID: 37034225
- PMCID: PMC10077036
PTPMT1 regulates mitochondrial death through the SLC25A6-NDUFS2 axis in pancreatic cancer cells
Abstract
Pancreatic ductal adenocarcinoma is a highly malignant cancer with poor prognosis, for which effective therapeutic strategies are urgently needed. The dual-specificity phosphatase PTPMT1 is localized in mitochondria and highly expressed in various cancers. Here, we investigated the function of PTPMT1 in pancreatic ductal adenocarcinoma. We inhibited its expression in pancreatic cancer cell lines using siRNAs or the specific PTPMT1 inhibitor alexidine dihydrochloride and observed that PTPMT1 silencing in pancreatic cancer cell lines drastically reduced cell viability, caused mitochondrial damage, and impaired mitochondrial function. Co-immunoprecipitation analysis demonstrated that PTPMT1 could interact with SLC25A6 and NDUFS2, indicating that it may modulate mitochondrial function via the SLC25A6-NDUFS2 axis. Collecively, our data highlight PTPMT1 as an important factor in pancreatic ductal adenocarcinoma and a potential therapeutic target.
Keywords: PTPMT1; Pancreatic ductal adenocarcinoma; SLC25A6; alexidine dihydrochloride.
AJCR Copyright © 2023.
Conflict of interest statement
None.
Figures
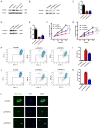




Similar articles
-
PTPMT1 inhibition induces apoptosis and growth arrest of human SCLC cells by disrupting mitochondrial metabolism.Transl Cancer Res. 2024 Dec 31;13(12):6956-6969. doi: 10.21037/tcr-2024-2379. Epub 2024 Dec 27. Transl Cancer Res. 2024. PMID: 39816544 Free PMC article.
-
Pharmacological targeting of the mitochondrial phosphatase PTPMT1.J Pharmacol Exp Ther. 2010 May;333(2):584-92. doi: 10.1124/jpet.109.163329. Epub 2010 Feb 18. J Pharmacol Exp Ther. 2010. PMID: 20167843 Free PMC article.
-
Downregulation of the mitochondrial phosphatase PTPMT1 is sufficient to promote cancer cell death.PLoS One. 2013;8(1):e53803. doi: 10.1371/journal.pone.0053803. Epub 2013 Jan 10. PLoS One. 2013. PMID: 23326511 Free PMC article.
-
Metformin as an Adjunctive Therapy for Pancreatic Cancer: A Review of the Literature on Its Potential Therapeutic Use.Dig Dis Sci. 2018 Nov;63(11):2840-2852. doi: 10.1007/s10620-018-5233-y. Epub 2018 Aug 29. Dig Dis Sci. 2018. PMID: 30159732 Review.
-
Higher notch expression implies poor survival in pancreatic ductal adenocarcinoma: A systematic review and meta-analysis.Pancreatology. 2018 Dec;18(8):954-961. doi: 10.1016/j.pan.2018.09.014. Epub 2018 Oct 1. Pancreatology. 2018. PMID: 30297095
Cited by
-
OTUB1/NDUFS2 axis promotes pancreatic tumorigenesis through protecting against mitochondrial cell death.Cell Death Discov. 2024 Apr 23;10(1):190. doi: 10.1038/s41420-024-01948-x. Cell Death Discov. 2024. PMID: 38653740 Free PMC article.
-
NFκB1: a common biomarker linking Alzheimer's and Parkinson's disease pathology.Front Neurosci. 2025 May 6;19:1589857. doi: 10.3389/fnins.2025.1589857. eCollection 2025. Front Neurosci. 2025. PMID: 40395692 Free PMC article.
-
PTPMT1 inhibition induces apoptosis and growth arrest of human SCLC cells by disrupting mitochondrial metabolism.Transl Cancer Res. 2024 Dec 31;13(12):6956-6969. doi: 10.21037/tcr-2024-2379. Epub 2024 Dec 27. Transl Cancer Res. 2024. PMID: 39816544 Free PMC article.
-
A novel mitochondrial-related risk model for predicting prognosis and immune checkpoint blockade therapy response in uterine corpus endometrial carcinoma.Sci Rep. 2025 Jan 9;15(1):1404. doi: 10.1038/s41598-025-85537-7. Sci Rep. 2025. PMID: 39789247 Free PMC article.
-
Pharmacological targeting of the mitochondrial phosphatase PTPMT1 sensitizes hepatocellular carcinoma to ferroptosis.Cell Death Dis. 2025 Apr 6;16(1):257. doi: 10.1038/s41419-025-07581-5. Cell Death Dis. 2025. PMID: 40189563 Free PMC article.
References
-
- Advancing on pancreatic cancer. Nat Rev Gastroenterol Hepatol. 2021;18:447. - PubMed
-
- Jain T, Dudeja V. The war against pancreatic cancer in 2020-advances on all fronts. Nat Rev Gastroenterol Hepatol. 2021;18:99–100. - PubMed
-
- Pagliarini DJ, Worby CA, Dixon JE. A PTEN-like phosphatase with a novel substrate specificity. J Biol Chem. 2004;279:38590–38596. - PubMed
LinkOut - more resources
Full Text Sources
