This is a preprint.
A single neuron in C. elegans orchestrates multiple motor outputs through parallel modes of transmission
- PMID: 37034579
- PMCID: PMC10081309
- DOI: 10.1101/2023.04.02.532814
A single neuron in C. elegans orchestrates multiple motor outputs through parallel modes of transmission
Update in
-
A single neuron in C. elegans orchestrates multiple motor outputs through parallel modes of transmission.Curr Biol. 2023 Oct 23;33(20):4430-4445.e6. doi: 10.1016/j.cub.2023.08.088. Epub 2023 Sep 27. Curr Biol. 2023. PMID: 37769660 Free PMC article.
Abstract
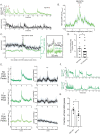
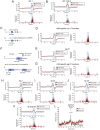
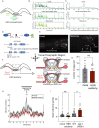
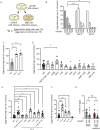
Animals generate a wide range of highly coordinated motor outputs, which allows them to execute purposeful behaviors. Individual neuron classes in the circuits that generate behavior have a remarkable capacity for flexibility, as they exhibit multiple axonal projections, transmitter systems, and modes of neural activity. How these multi-functional properties of neurons enable the generation of highly coordinated behaviors remains unknown. Here we show that the HSN neuron in C. elegans evokes multiple motor programs over different timescales to enable a suite of behavioral changes during egg-laying. Using HSN activity perturbations and in vivo calcium imaging, we show that HSN acutely increases egg-laying and locomotion while also biasing the animals towards low-speed dwelling behavior over longer timescales. The acute effects of HSN on egg-laying and high-speed locomotion are mediated by separate sets of HSN transmitters and different HSN axonal projections. The long-lasting effects on dwelling are mediated by HSN release of serotonin that is taken up and re-released by NSM, another serotonergic neuron class that directly evokes dwelling. Our results show how the multi-functional properties of a single neuron allow it to induce a coordinated suite of behaviors and also reveal for the first time that neurons can borrow serotonin from one another to control behavior.
Conflict of interest statement
DECLARATION OF INTERESTS
The authors have no competing interests to declare.
Figures


References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
