Development and evaluation of a java-based deep neural network method for drug response predictions
- PMID: 37035534
- PMCID: PMC10076891
- DOI: 10.3389/frai.2023.1069353
Development and evaluation of a java-based deep neural network method for drug response predictions
Abstract
Accurate prediction of drug response is a crucial step in personalized medicine. Recently, deep learning techniques have been witnessed with significant breakthroughs in a variety of areas including biomedical research and chemogenomic applications. This motivated us to develop a novel deep learning platform to accurately and reliably predict the response of cancer cells to different drug treatments. In the present work, we describe a Java-based implementation of deep neural network method, termed JavaDL, to predict cancer responses to drugs solely based on their chemical features. To this end, we devised a novel cost function and added a regularization term which suppresses overfitting. We also adopted an early stopping strategy to further reduce overfit and improve the accuracy and robustness of our models. To evaluate our method, we compared with several popular machine learning and deep neural network programs and observed that JavaDL either outperformed those methods in model building or obtained comparable predictions. Finally, JavaDL was employed to predict drug responses of several aggressive breast cancer cell lines, and the results showed robust and accurate predictions with r 2 as high as 0.81.
Keywords: artificial intelligence (AI); deep learning; deep neural network; drug response; multilayer neural network (MNN); quantitative structure activity relationship (QSAR); triple-negative breast cancer (TNBC).
Copyright © 2023 Huang, Fong, Chaudhari and Zhang.
Conflict of interest statement
Author RC is currently employed by the company Eurofins Beacon Discovery. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
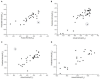
Figures

Similar articles
-
The rise of deep learning and transformations in bioactivity prediction power of molecular modeling tools.Chem Biol Drug Des. 2021 Nov;98(5):954-967. doi: 10.1111/cbdd.13750. Epub 2021 Sep 16. Chem Biol Drug Des. 2021. PMID: 34532977
-
Data Integration Using Advances in Machine Learning in Drug Discovery and Molecular Biology.Methods Mol Biol. 2021;2190:167-184. doi: 10.1007/978-1-0716-0826-5_7. Methods Mol Biol. 2021. PMID: 32804365 Review.
-
Artificial Intelligence-Based Traditional Chinese Medicine Assistive Diagnostic System: Validation Study.JMIR Med Inform. 2020 Jun 15;8(6):e17608. doi: 10.2196/17608. JMIR Med Inform. 2020. PMID: 32538797 Free PMC article.
-
Dissecting Machine-Learning Prediction of Molecular Activity: Is an Applicability Domain Needed for Quantitative Structure-Activity Relationship Models Based on Deep Neural Networks?J Chem Inf Model. 2019 Jan 28;59(1):117-126. doi: 10.1021/acs.jcim.8b00348. Epub 2018 Nov 21. J Chem Inf Model. 2019. PMID: 30412667
-
Role of Artificial Intelligence and Machine Learning in Prediction, Diagnosis, and Prognosis of Cancer.Cureus. 2022 Nov 2;14(11):e31008. doi: 10.7759/cureus.31008. eCollection 2022 Nov. Cureus. 2022. PMID: 36475188 Free PMC article. Review.
Cited by
-
Developing and validating a drug recommendation system based on tumor microenvironment and drug fingerprint.Front Artif Intell. 2025 Jan 8;7:1444127. doi: 10.3389/frai.2024.1444127. eCollection 2024. Front Artif Intell. 2025. PMID: 39850847 Free PMC article.
-
Optimized models and deep learning methods for drug response prediction in cancer treatments: a review.PeerJ Comput Sci. 2024 Mar 25;10:e1903. doi: 10.7717/peerj-cs.1903. eCollection 2024. PeerJ Comput Sci. 2024. PMID: 38660174 Free PMC article.
-
Application of Deep Learning in Cancer Prognosis Prediction Model.Technol Cancer Res Treat. 2023 Jan-Dec;22:15330338231199287. doi: 10.1177/15330338231199287. Technol Cancer Res Treat. 2023. PMID: 37709267 Free PMC article.
References
-
- Ballabio D., Manganaro A., Consonni V., Mauri A. (2009). Introduction to MOLE DB - on-line molecular descriptors database. Match Commun. Math. Comput. Chemist. 62, 199–207.
LinkOut - more resources
Full Text Sources

