Landscape of prostate-specific membrane antigen heterogeneity and regulation in AR-positive and AR-negative metastatic prostate cancer
- PMID: 37038004
- PMCID: PMC10867901
- DOI: 10.1038/s43018-023-00539-6
Landscape of prostate-specific membrane antigen heterogeneity and regulation in AR-positive and AR-negative metastatic prostate cancer
Abstract
Tumor expression of prostate-specific membrane antigen (PSMA) is lost in 15-20% of men with castration-resistant prostate cancer (CRPC), yet the underlying mechanisms remain poorly defined. In androgen receptor (AR)-positive CRPC, we observed lower PSMA expression in liver lesions versus other sites, suggesting a role of the microenvironment in modulating PSMA. PSMA suppression was associated with promoter histone 3 lysine 27 methylation and higher levels of neutral amino acid transporters, correlating with 18F-fluciclovine uptake on positron emission tomography imaging. While PSMA is regulated by AR, we identified a subset of AR-negative CRPC with high PSMA. HOXB13 and AR co-occupancy at the PSMA enhancer and knockout models point to HOXB13 as an upstream regulator of PSMA in AR-positive and AR-negative prostate cancer. These data demonstrate how PSMA expression is differentially regulated across metastatic lesions and in the context of the AR, which may inform selection for PSMA-targeted therapies and development of complementary biomarkers.
© 2023. The Author(s), under exclusive licence to Springer Nature America, Inc.
Conflict of interest statement
Competing interests
All other authors have no competing interests.
Figures








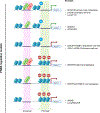






References
-
- Israeli RS, Powell CT, Corr JG, Fair WR & Heston WD Expression of the prostate-specific membrane antigen. Cancer Res. 54, 1807–1811 (1994). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

