Pten inhibition dedifferentiates long-distance axon-regenerating intrinsically photosensitive retinal ganglion cells and upregulates mitochondria-associated Dynlt1a and Lars2
- PMID: 37039265
- PMCID: PMC10163351
- DOI: 10.1242/dev.201644
Pten inhibition dedifferentiates long-distance axon-regenerating intrinsically photosensitive retinal ganglion cells and upregulates mitochondria-associated Dynlt1a and Lars2
Abstract
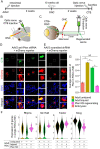
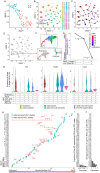
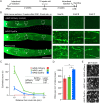
Central nervous system projection neurons fail to spontaneously regenerate injured axons. Targeting developmentally regulated genes in order to reactivate embryonic intrinsic axon growth capacity or targeting pro-growth tumor suppressor genes such as Pten promotes long-distance axon regeneration in only a small subset of injured retinal ganglion cells (RGCs), despite many RGCs regenerating short-distance axons. A recent study identified αRGCs as the primary type that regenerates short-distance axons in response to Pten inhibition, but the rare types which regenerate long-distance axons, and cellular features that enable such response, remained unknown. Here, we used a new method for capturing specifically the rare long-distance axon-regenerating RGCs, and also compared their transcriptomes with embryonic RGCs, in order to answer these questions. We found the existence of adult non-α intrinsically photosensitive M1 RGC subtypes that retained features of embryonic cell state, and showed that these subtypes partially dedifferentiated towards an embryonic state and regenerated long-distance axons in response to Pten inhibition. We also identified Pten inhibition-upregulated mitochondria-associated genes, Dynlt1a and Lars2, which promote axon regeneration on their own, and thus present novel therapeutic targets.
Keywords: Axon growth; Mouse; Optic nerve; Retinal ganglion cell; scRNA-seq.
© 2023. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interests The authors declare no competing or financial interests.
Figures


References
-
- Blackmore, M. G., Wang, Z., Lerch, J. K., Motti, D., Zhang, Y. P., Shields, C. B., Lee, J. K., Goldberg, J. L., Lemmon, V. P. and Bixby, J. L. (2012). Krüppel-like Factor 7 engineered for transcriptional activation promotes axon regeneration in the adult corticospinal tract. Proc. Natl. Acad. Sci. USA 109, 7517-7522. 10.1073/pnas.1120684109 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

