The small compound Icerguastat reduces muscle defects in oculopharyngeal muscular dystrophy through the PERK pathway of the unfolded protein response
- PMID: 37042114
- PMCID: PMC10090878
- DOI: 10.1098/rsob.230008
The small compound Icerguastat reduces muscle defects in oculopharyngeal muscular dystrophy through the PERK pathway of the unfolded protein response
Abstract
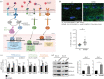
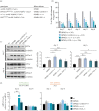
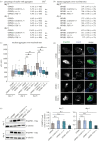
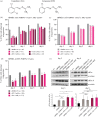
Oculopharyngeal muscular dystrophy (OPMD) is an autosomal dominant disease characterized by the progressive degeneration of specific muscles. OPMD is due to a mutation in the gene encoding poly(A) binding protein nuclear 1 (PABPN1) leading to a stretch of 11 to 18 alanines at N-terminus of the protein, instead of 10 alanines in the normal protein. This alanine tract extension induces the misfolding and aggregation of PABPN1 in muscle nuclei. Here, using Drosophila OPMD models, we show that the unfolded protein response (UPR) is activated in OPMD upon endoplasmic reticulum stress. Mutations in components of the PERK branch of the UPR reduce muscle degeneration and PABPN1 aggregation characteristic of the disease. We show that oral treatment of OPMD flies with Icerguastat (previously IFB-088), a Guanabenz acetate derivative that shows lower side effects, also decreases muscle degeneration and PABPN1 aggregation. Furthermore, the positive effect of Icerguastat depends on GADD34, a key component of the phosphatase complex in the PERK branch of the UPR. This study reveals a major contribution of the ER stress in OPMD pathogenesis and provides a proof-of-concept for Icerguastat interest in future pharmacological treatments of OPMD.
Keywords: Drosophila model; GADD34; Icerguastat/IFB-088; OPMD; PEK/PERK; unfolded protein response.
Conflict of interest statement
E.A. is a full-time employee of InFlectis BioScience and P.G. is a full-time employee and founder of InFlectis BioScience.
Figures


Similar articles
-
Activation of the ubiquitin-proteasome system contributes to oculopharyngeal muscular dystrophy through muscle atrophy.PLoS Genet. 2022 Jan 13;18(1):e1010015. doi: 10.1371/journal.pgen.1010015. eCollection 2022 Jan. PLoS Genet. 2022. PMID: 35025870 Free PMC article.
-
Functional impact of an oculopharyngeal muscular dystrophy mutation in PABPN1.J Physiol. 2017 Jul 1;595(13):4167-4187. doi: 10.1113/JP273948. Epub 2017 Apr 25. J Physiol. 2017. PMID: 28303574 Free PMC article.
-
A Drosophila model of oculopharyngeal muscular dystrophy reveals intrinsic toxicity of PABPN1.EMBO J. 2006 May 17;25(10):2253-62. doi: 10.1038/sj.emboj.7601117. Epub 2006 Apr 27. EMBO J. 2006. PMID: 16642034 Free PMC article.
-
Progress in understanding the pathogenesis of oculopharyngeal muscular dystrophy.Can J Neurol Sci. 2003 Feb;30(1):8-14. doi: 10.1017/s0317167100002365. Can J Neurol Sci. 2003. PMID: 12619777 Review.
-
Oculopharyngeal muscular dystrophy: recent advances in the understanding of the molecular pathogenic mechanisms and treatment strategies.Biochim Biophys Acta. 2007 Feb;1772(2):173-85. doi: 10.1016/j.bbadis.2006.10.003. Epub 2006 Oct 11. Biochim Biophys Acta. 2007. PMID: 17110089 Review.
Cited by
-
Drosophila melanogaster: A Model Organism in Muscular Dystrophy Studies.Int J Mol Sci. 2025 Feb 10;26(4):1459. doi: 10.3390/ijms26041459. Int J Mol Sci. 2025. PMID: 40003927 Free PMC article. Review.
-
Decoding Nucleotide Repeat Expansion Diseases: Novel Insights from Drosophila melanogaster Studies.Int J Mol Sci. 2024 Nov 2;25(21):11794. doi: 10.3390/ijms252111794. Int J Mol Sci. 2024. PMID: 39519345 Free PMC article. Review.
-
Aligning with the 3Rs: alternative models for research into muscle development and inherited myopathies.BMC Vet Res. 2024 Oct 18;20(1):477. doi: 10.1186/s12917-024-04309-z. BMC Vet Res. 2024. PMID: 39425123 Free PMC article. Review.
-
Aryl Guanyl Hydrazones: A Viable Strategy for Designing BBB-Permeable, Neuroactive Compounds?ACS Chem Neurosci. 2025 Aug 6;16(15):2767-2775. doi: 10.1021/acschemneuro.5c00463. Epub 2025 Jul 22. ACS Chem Neurosci. 2025. PMID: 40693920 Free PMC article. Review.
References
Publication types
MeSH terms
Associated data
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

