Investigation of the Entry Pathway and Molecular Nature of σ1 Receptor Ligands
- PMID: 37047338
- PMCID: PMC10094450
- DOI: 10.3390/ijms24076367
Investigation of the Entry Pathway and Molecular Nature of σ1 Receptor Ligands
Abstract
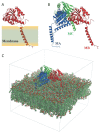
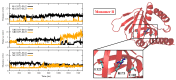
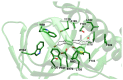
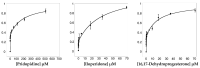
The σ1 receptor (σ1-R) is an enigmatic endoplasmic reticulum resident transmembrane protein implicated in a variety of central nervous system disorders and whose agonists have neuroprotective activity. In spite of σ1-R's physio-pathological and pharmacological importance, two of the most important features required to fully understand σ1-R function, namely the receptor endogenous ligand(s) and the molecular mechanism of ligand access to the binding site, have not yet been unequivocally determined. In this work, we performed molecular dynamics (MD) simulations to help clarify the potential route of access of ligand(s) to the σ1-R binding site, on which discordant results had been reported in the literature. Further, we combined computational and experimental procedures (i.e., virtual screening (VS), electron density map fitting and fluorescence titration experiments) to provide indications about the nature of σ1-R endogenous ligand(s). Our MD simulations on human σ1-R suggested that ligands access the binding site through a cavity that opens on the protein surface in contact with the membrane, in agreement with previous experimental studies on σ1-R from Xenopus laevis. Additionally, steroids were found to be among the preferred σ1-R ligands predicted by VS, and 16,17-didehydroprogesterone was shown by fluorescence titration to bind human σ1-R, with significantly higher affinity than the prototypic σ1-R ligand pridopidine in the same essay. These results support the hypothesis that steroids are among the most important physiological σ1-R ligands.
Keywords: Huntington’s disease; fluorescence titration; molecular dynamics; virtual screening; σ1 receptor.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
The sigma enigma: in vitro/in silico site-directed mutagenesis studies unveil σ1 receptor ligand binding.Biochemistry. 2014 May 13;53(18):2993-3003. doi: 10.1021/bi401575g. Epub 2014 Apr 28. Biochemistry. 2014. PMID: 24766040
-
Chemoenzymatic synthesis of 2,6-disubstituted tetrahydropyrans with high σ1 receptor affinity, antitumor and analgesic activity.Eur J Med Chem. 2021 Jul 5;219:113443. doi: 10.1016/j.ejmech.2021.113443. Epub 2021 Apr 20. Eur J Med Chem. 2021. PMID: 33901806
-
New Insights into the Opening of the Occluded Ligand-Binding Pocket of Sigma1 Receptor: Binding of a Novel Bivalent RC-33 Derivative.J Chem Inf Model. 2020 Feb 24;60(2):756-765. doi: 10.1021/acs.jcim.9b00649. Epub 2019 Dec 20. J Chem Inf Model. 2020. PMID: 31809025
-
Structural Insights into Sigma1 Function.Handb Exp Pharmacol. 2017;244:13-25. doi: 10.1007/164_2016_95. Handb Exp Pharmacol. 2017. PMID: 27995388 Review.
-
Medicinal Chemistry of σ1 Receptor Ligands: Pharmacophore Models, Synthesis, Structure Affinity Relationships, and Pharmacological Applications.Handb Exp Pharmacol. 2017;244:51-79. doi: 10.1007/164_2017_33. Handb Exp Pharmacol. 2017. PMID: 28620761 Review.
Cited by
-
Insight into binding of endogenous neurosteroid ligands to the sigma-1 receptor.Nat Commun. 2024 Jul 4;15(1):5619. doi: 10.1038/s41467-024-49894-7. Nat Commun. 2024. PMID: 38965213 Free PMC article.
-
Comprehensive Analysis of Age- and Sex-Related Expression of the Chaperone Protein Sigma-1R in the Mouse Brain.Brain Sci. 2024 Aug 30;14(9):881. doi: 10.3390/brainsci14090881. Brain Sci. 2024. PMID: 39335377 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

