Comparative Transcriptome Analysis Reveals a Potential Regulatory Network for Ogura Cytoplasmic Male Sterility in Cabbage (Brassica oleracea L.)
- PMID: 37047676
- PMCID: PMC10094764
- DOI: 10.3390/ijms24076703
Comparative Transcriptome Analysis Reveals a Potential Regulatory Network for Ogura Cytoplasmic Male Sterility in Cabbage (Brassica oleracea L.)
Abstract
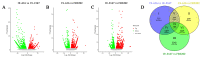
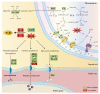
Ogura cytoplasmic male sterility (CMS) lines are widely used breeding materials in cruciferous crops and play important roles in heterosis utilization; however, the sterility mechanism remains unclear. To investigate the microspore development process and gene expression changes after the introduction of orf138 and Rfo, cytological observation and transcriptome analysis were performed using a maintainer line, an Ogura CMS line, and a restorer line. Semithin sections of microspores at different developmental stages showed that the degradation of tapetal cells began at the tetrad stage in the Ogura CMS line, while it occurred at the bicellular microspore stage to the tricellular microspore stage in the maintainer and restorer lines. Therefore, early degradation of tapetal cells may be the cause of pollen abortion. Transcriptome analysis results showed that a total of 1287 DEGs had consistent expression trends in the maintainer line and restorer line, but were significantly up- or down-regulated in the Ogura CMS line, indicating that they may be closely related to pollen abortion. Functional annotation showed that the 1287 core DEGs included a large number of genes related to pollen development, oxidative phosphorylation, carbohydrate, lipid, and protein metabolism. In addition, further verification elucidated that down-regulated expression of genes related to energy metabolism led to decreased ATP content and excessive ROS accumulation in the anthers of Ogura CMS. Based on these results, we propose a transcriptome-mediated induction and regulatory network for cabbage Ogura CMS. Our research provides new insights into the mechanism of pollen abortion and fertility restoration in Ogura CMS.
Keywords: Ogura CMS; cabbage; comparative transcriptome analysis; pollen development.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




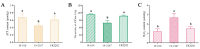
Similar articles
-
Integrated Transcriptomics and Metabolomics Analysis Reveals Convergent and Divergent Key Molecular Networks of Dominant Genic Male Sterility and Cytoplasmic Male Sterility in Cabbage.Int J Mol Sci. 2025 Jan 31;26(3):1259. doi: 10.3390/ijms26031259. Int J Mol Sci. 2025. PMID: 39941029 Free PMC article.
-
Comparative analysis of the complete mitochondrial genome sequences and anther development cytology between maintainer and Ogura-type cytoplasm male-sterile cabbage (B. oleracea Var. capitata).BMC Genomics. 2021 Sep 7;22(1):646. doi: 10.1186/s12864-021-07963-x. BMC Genomics. 2021. PMID: 34493212 Free PMC article.
-
Ogura-CMS in Chinese cabbage (Brassica rapa ssp. pekinensis) causes delayed expression of many nuclear genes.Plant Sci. 2013 Feb;199-200:7-17. doi: 10.1016/j.plantsci.2012.11.001. Epub 2012 Nov 12. Plant Sci. 2013. PMID: 23265314
-
Mechanism and Utilization of Ogura Cytoplasmic Male Sterility in Cruciferae Crops.Int J Mol Sci. 2022 Aug 13;23(16):9099. doi: 10.3390/ijms23169099. Int J Mol Sci. 2022. PMID: 36012365 Free PMC article. Review.
-
Past and future of cytoplasmic male sterility and heterosis breeding in crop plants.Plant Cell Rep. 2025 Jan 22;44(2):33. doi: 10.1007/s00299-024-03414-5. Plant Cell Rep. 2025. PMID: 39841239 Review.
Cited by
-
Defense strategies and associated phytohormonal regulation in Brassica plants in response to chewing and sap-sucking insects.Front Plant Sci. 2024 Apr 5;15:1376917. doi: 10.3389/fpls.2024.1376917. eCollection 2024. Front Plant Sci. 2024. PMID: 38645389 Free PMC article. Review.
-
Integrated Cytological, Physiological, and Comparative Transcriptome Profiling Analysis of the Male Sterility Mechanism of 'Xinli No.7' Pear (Pyrus sp.).Plants (Basel). 2025 Jun 11;14(12):1783. doi: 10.3390/plants14121783. Plants (Basel). 2025. PMID: 40573771 Free PMC article.
References
-
- FAO Food and Agriculture Organization of the United Nations, FAOSTAT. 2021. [(accessed on 25 December 2022)]. Available online: https://www.fao.org/faostat/zh/#data/QCL.
-
- Kuera V., Věra C., Miroslava V., Miroslav K. Hybrid breeding of cauliflower using self-incompatibility and cytoplasmic male sterility. Hortic. Sci. (HORTSCI) 2006;33:148–152. doi: 10.17221/3754-HORTSCI. - DOI
-
- Dey S.S., Singh N., Bhatia R., Parkash C., Chandel C. Genetic combining ability and heterosis for important vitamins and antioxidant pigments in cauliflower (Brassica oleracea var. botrytis L.) Euphytica. 2014;195:169–181. doi: 10.1007/s10681-013-0981-4. - DOI
-
- Singh S., Dey S.S., Bhatia R., Kumar R., Sharma K., Behera T.K. Heterosis and combining ability in cytoplasmic male sterile and doubled haploid based Brassica oleracea progenies and prediction of heterosis using microsatellites. PLoS ONE. 2019;14:e0210772. doi: 10.1371/journal.pone.0210772. - DOI - PMC - PubMed
-
- Ren W., Li Z., Han F., Zhang B., Li X., Fang Z., Yang L., Zhuang M., Lv H., Liu Y., et al. Utilization of Ogura CMS germplasm with the clubroot resistance gene by fertility restoration and cytoplasm replacement in Brassica oleracea L. Hortic. Res. 2020;7:61. doi: 10.1038/s41438-020-0282-8. - DOI - PMC - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources

