Salt-Specific Suppression of the Cold Denaturation of Thermophilic Multidomain Initiation Factor 2
- PMID: 37047761
- PMCID: PMC10094840
- DOI: 10.3390/ijms24076787
Salt-Specific Suppression of the Cold Denaturation of Thermophilic Multidomain Initiation Factor 2
Abstract
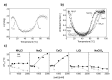
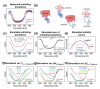
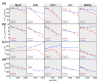
Thermophilic proteins and enzymes are attractive for use in industrial applications due to their resistance against heat and denaturants. Here, we report on a thermophilic protein that is stable at high temperatures (Ttrs, hot 67 °C) but undergoes significant unfolding at room temperature due to cold denaturation. Little is known about the cold denaturation of thermophilic proteins, although it can significantly limit their applications. We investigated the cold denaturation of thermophilic multidomain protein translation initiation factor 2 (IF2) from Thermus thermophilus. IF2 is a GTPase that binds to ribosomal subunits and initiator fMet-tRNAfMet during the initiation of protein biosynthesis. In the presence of 9 M urea, measurements in the far-UV region by circular dichroism were used to capture details about the secondary structure of full-length IF2 protein and its domains during cold and hot denaturation. Cold denaturation can be suppressed by salt, depending on the type, due to the decreased heat capacity. Thermodynamic analysis and mathematical modeling of the denaturation process showed that salts reduce the cooperativity of denaturation of the IF2 domains, which might be associated with the high frustration between domains. This characteristic of high interdomain frustration may be the key to satisfying numerous diverse contacts with ribosomal subunits, translation factors, and tRNA.
Keywords: cold denaturation; domain cooperativity; electrostatic interactions; initiation factor 2; thermodynamic stability.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
-
- Jaenicke R., Böhm G. Methods in Enzymology. Academic Press; Cambridge, MA, USA: 2001. [33] Thermostability of Proteins from Thermotoga Maritima; pp. 438–469. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

