KDEL Receptor Trafficking to the Plasma Membrane Is Regulated by ACBD3 and Rab4A-GTP
- PMID: 37048152
- PMCID: PMC10093020
- DOI: 10.3390/cells12071079
KDEL Receptor Trafficking to the Plasma Membrane Is Regulated by ACBD3 and Rab4A-GTP
Abstract
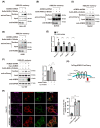
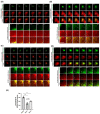
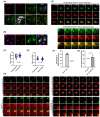
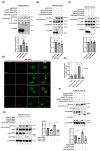
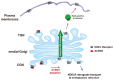
KDEL receptor-1 maintains homeostasis in the early secretory pathway by capturing and retrieving ER chaperones to the ER during heavy secretory activity. Unexpectedly, a fraction of the receptor is also known to reside in the plasma membrane (PM), although it is largely unknown exactly how the KDEL receptor gets exported from the Golgi and travels to the PM. We have previously shown that a Golgi scaffolding protein (ACBD3) facilitates KDEL receptor localization at the Golgi via the regulating cargo wave-induced cAMP/PKA-dependent signaling pathway. Upon endocytosis, surface-expressed KDEL receptor undergoes highly complex itineraries through the Golgi and the endo-lysosomal compartments, where the endocytosed receptor utilizes Rab14A- and Rab11A-positive recycling endosomes and clathrin-decorated tubulovesicular carriers. In this study, we sought to investigate the mechanism through which the KDEL receptor gets exported from the Golgi en route to the PM. We report here that ACBD3 depletion results in greatly increased trafficking of KDEL receptor to the PM via Rab4A-positive tubular carriers emanating from the Golgi. Expression of constitutively activated Rab4A mutant (Q72L) increases the surface expression of KDEL receptor up to 2~3-fold, whereas Rab4A knockdown or the expression of GDP-locked Rab4A mutant (S27N) inhibits KDEL receptor targeting of the PM. Importantly, KDELR trafficking from the Golgi to the PM is independent of PKA- and Src kinase-mediated mechanisms. Taken together, these results reveal that ACBD3 and Rab4A play a key role in regulating KDEL receptor trafficking to the cell surface.
Keywords: ACBD3; Golgi; KDEL receptor; Rab11A; Rab4A; plasma membrane.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
ACBD3 modulates KDEL receptor interaction with PKA for its trafficking via tubulovesicular carrier.BMC Biol. 2021 Sep 7;19(1):194. doi: 10.1186/s12915-021-01137-7. BMC Biol. 2021. PMID: 34493279 Free PMC article.
-
KDEL receptor is a cell surface receptor that cycles between the plasma membrane and the Golgi via clathrin-mediated transport carriers.Cell Mol Life Sci. 2021 Feb;78(3):1085-1100. doi: 10.1007/s00018-020-03570-3. Epub 2020 Jun 19. Cell Mol Life Sci. 2021. PMID: 32562023 Free PMC article.
-
An A-kinase anchoring protein (ACBD3) coordinates traffic-induced PKA activation at the Golgi.J Biol Chem. 2023 May;299(5):104696. doi: 10.1016/j.jbc.2023.104696. Epub 2023 Apr 10. J Biol Chem. 2023. PMID: 37044218 Free PMC article.
-
Regulation of Golgi signaling and trafficking by the KDEL receptor.Histochem Cell Biol. 2013 Oct;140(4):395-405. doi: 10.1007/s00418-013-1130-9. Epub 2013 Jul 20. Histochem Cell Biol. 2013. PMID: 23873287 Review.
-
Molecular basis for KDEL-mediated retrieval of escaped ER-resident proteins - SWEET talking the COPs.J Cell Sci. 2020 Oct 9;133(19):jcs250100. doi: 10.1242/jcs.250100. J Cell Sci. 2020. PMID: 33037041 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

