High-Throughput Microscopy Analysis of Mitochondrial Membrane Potential in 2D and 3D Models
- PMID: 37048162
- PMCID: PMC10093082
- DOI: 10.3390/cells12071089
High-Throughput Microscopy Analysis of Mitochondrial Membrane Potential in 2D and 3D Models
Abstract
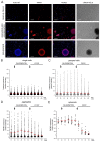
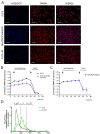
Recent proteomic, metabolomic, and transcriptomic studies have highlighted a connection between changes in mitochondria physiology and cellular pathophysiological mechanisms. Secondary assays to assess the function of these organelles appear fundamental to validate these -omics findings. Although mitochondrial membrane potential is widely recognized as an indicator of mitochondrial activity, high-content imaging-based approaches coupled to multiparametric to measure it have not been established yet. In this paper, we describe a methodology for the unbiased high-throughput quantification of mitochondrial membrane potential in vitro, which is suitable for 2D to 3D models. We successfully used our method to analyze mitochondrial membrane potential in monolayers of human fibroblasts, neural stem cells, spheroids, and isolated muscle fibers. Moreover, by combining automated image analysis and machine learning, we were able to discriminate melanoma cells from macrophages in co-culture and to analyze the subpopulations separately. Our data demonstrated that our method is a widely applicable strategy for large-scale profiling of mitochondrial activity.
Keywords: NSCs; TMRM; co-culture; machine learning; mitochondrial membrane potential; single muscle fibers; spheroids.
Conflict of interest statement
A.P.M.R. and M.G. are guest editors of the Special Issue.
Figures


References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

