Extracting relevant predictive variables for COVID-19 severity prognosis: An exhaustive comparison of feature selection techniques
- PMID: 37053151
- PMCID: PMC10101453
- DOI: 10.1371/journal.pone.0284150
Extracting relevant predictive variables for COVID-19 severity prognosis: An exhaustive comparison of feature selection techniques
Abstract
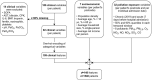
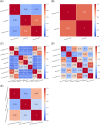
With the COVID-19 pandemic having caused unprecedented numbers of infections and deaths, large research efforts have been undertaken to increase our understanding of the disease and the factors which determine diverse clinical evolutions. Here we focused on a fully data-driven exploration regarding which factors (clinical or otherwise) were most informative for SARS-CoV-2 pneumonia severity prediction via machine learning (ML). In particular, feature selection techniques (FS), designed to reduce the dimensionality of data, allowed us to characterize which of our variables were the most useful for ML prognosis. We conducted a multi-centre clinical study, enrolling n = 1548 patients hospitalized due to SARS-CoV-2 pneumonia: where 792, 238, and 598 patients experienced low, medium and high-severity evolutions, respectively. Up to 106 patient-specific clinical variables were collected at admission, although 14 of them had to be discarded for containing ⩾60% missing values. Alongside 7 socioeconomic attributes and 32 exposures to air pollution (chronic and acute), these became d = 148 features after variable encoding. We addressed this ordinal classification problem both as a ML classification and regression task. Two imputation techniques for missing data were explored, along with a total of 166 unique FS algorithm configurations: 46 filters, 100 wrappers and 20 embeddeds. Of these, 21 setups achieved satisfactory bootstrap stability (⩾0.70) with reasonable computation times: 16 filters, 2 wrappers, and 3 embeddeds. The subsets of features selected by each technique showed modest Jaccard similarities across them. However, they consistently pointed out the importance of certain explanatory variables. Namely: patient's C-reactive protein (CRP), pneumonia severity index (PSI), respiratory rate (RR) and oxygen levels -saturation Sp O2, quotients Sp O2/RR and arterial Sat O2/Fi O2-, the neutrophil-to-lymphocyte ratio (NLR) -to certain extent, also neutrophil and lymphocyte counts separately-, lactate dehydrogenase (LDH), and procalcitonin (PCT) levels in blood. A remarkable agreement has been found a posteriori between our strategy and independent clinical research works investigating risk factors for COVID-19 severity. Hence, these findings stress the suitability of this type of fully data-driven approaches for knowledge extraction, as a complementary to clinical perspectives.
Copyright: © 2023 Hayet-Otero et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



Similar articles
-
Safety and Efficacy of Imatinib for Hospitalized Adults with COVID-19: A structured summary of a study protocol for a randomised controlled trial.Trials. 2020 Oct 28;21(1):897. doi: 10.1186/s13063-020-04819-9. Trials. 2020. PMID: 33115543 Free PMC article.
-
Factors associated with death outcome in patients with severe coronavirus disease-19 (COVID-19): a case-control study.Int J Med Sci. 2020 May 18;17(9):1281-1292. doi: 10.7150/ijms.46614. eCollection 2020. Int J Med Sci. 2020. PMID: 32547323 Free PMC article.
-
Predictive role of clinical features in patients with coronavirus disease 2019 for severe disease.Zhong Nan Da Xue Xue Bao Yi Xue Ban. 2020 May 28;45(5):536-541. doi: 10.11817/j.issn.1672-7347.2020.200384. Zhong Nan Da Xue Xue Bao Yi Xue Ban. 2020. PMID: 32879103 Chinese, English.
-
Artificial intelligence in clinical care amidst COVID-19 pandemic: A systematic review.Comput Struct Biotechnol J. 2021;19:2833-2850. doi: 10.1016/j.csbj.2021.05.010. Epub 2021 May 7. Comput Struct Biotechnol J. 2021. PMID: 34025952 Free PMC article. Review.
-
Laboratory features of severe vs. non-severe COVID-19 patients in Asian populations: a systematic review and meta-analysis.Eur J Med Res. 2020 Aug 3;25(1):30. doi: 10.1186/s40001-020-00432-3. Eur J Med Res. 2020. PMID: 32746929 Free PMC article.
Cited by
-
Obtaining patient phenotypes in SARS-CoV-2 pneumonia, and their association with clinical severity and mortality.Pneumonia (Nathan). 2024 Jun 25;16(1):12. doi: 10.1186/s41479-024-00132-0. Pneumonia (Nathan). 2024. PMID: 38915125 Free PMC article.
-
Comparative analysis of feature selection techniques for COVID-19 dataset.Sci Rep. 2024 Aug 11;14(1):18627. doi: 10.1038/s41598-024-69209-6. Sci Rep. 2024. PMID: 39128991 Free PMC article.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

