Mapping CircRNA-miRNA-mRNA regulatory axis identifies hsa_circ_0080942 and hsa_circ_0080135 as a potential theranostic agents for SARS-CoV-2 infection
- PMID: 37053191
- PMCID: PMC10101458
- DOI: 10.1371/journal.pone.0283589
Mapping CircRNA-miRNA-mRNA regulatory axis identifies hsa_circ_0080942 and hsa_circ_0080135 as a potential theranostic agents for SARS-CoV-2 infection
Abstract
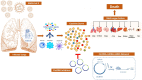
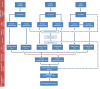
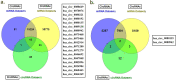
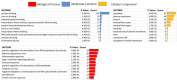
Non-coding RNAs (ncRNAs) can control the flux of genetic information; affect RNA stability and play crucial roles in mediating epigenetic modifications. A number of studies have highlighted the potential roles of both virus-encoded and host-encoded ncRNAs in viral infections, transmission and therapeutics. However, the role of an emerging type of non-coding transcript, circular RNA (circRNA) in severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection has not been fully elucidated so far. Moreover, the potential pathogenic role of circRNA-miRNA-mRNA regulatory axis has not been fully explored as yet. The current study aimed to holistically map the regulatory networks driven by SARS-CoV-2 related circRNAs, miRNAs and mRNAs to uncover plausible interactions and interplay amongst them in order to explore possible therapeutic options in SARS-CoV-2 infection. Patient datasets were analyzed systematically in a unified approach to explore circRNA, miRNA, and mRNA expression profiles. CircRNA-miRNA-mRNA network was constructed based on cytokine storm related circRNAs forming a total of 165 circRNA-miRNA-mRNA pairs. This study implies the potential regulatory role of the obtained circRNA-miRNA-mRNA network and proposes that two differentially expressed circRNAs hsa_circ_0080942 and hsa_circ_0080135 might serve as a potential theranostic agents for SARS-CoV-2 infection. Collectively, the results shed light on the functional role of circRNAs as ceRNAs to sponge miRNA and regulate mRNA expression during SARS-CoV-2 infection.
Copyright: © 2023 Ayaz et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
Construction of the circRNA-miRNA-mRNA Regulatory Network of an Abdominal Aortic Aneurysm to Explore Its Potential Pathogenesis.Dis Markers. 2021 Nov 5;2021:9916881. doi: 10.1155/2021/9916881. eCollection 2021. Dis Markers. 2021. PMID: 34777635 Free PMC article.
-
Construct a circRNA/miRNA/mRNA regulatory network to explore potential pathogenesis and therapy options of clear cell renal cell carcinoma.Sci Rep. 2020 Aug 12;10(1):13659. doi: 10.1038/s41598-020-70484-2. Sci Rep. 2020. PMID: 32788609 Free PMC article.
-
Competing endogenous RNA network mediated by circ_3205 in SARS-CoV-2 infected cells.Cell Mol Life Sci. 2022 Jan 17;79(2):75. doi: 10.1007/s00018-021-04119-8. Cell Mol Life Sci. 2022. PMID: 35039944 Free PMC article.
-
RNA-Seq Revealed a Circular RNA-microRNA-mRNA Regulatory Network in Hantaan Virus Infection.Front Cell Infect Microbiol. 2020 Mar 13;10:97. doi: 10.3389/fcimb.2020.00097. eCollection 2020. Front Cell Infect Microbiol. 2020. PMID: 32232013 Free PMC article. Review.
-
CircRNAs: a family number of miRNA regulatory transcriptome in laryngeal carcinoma.J Clin Lab Anal. 2021 Nov;35(11):e24038. doi: 10.1002/jcla.24038. Epub 2021 Oct 7. J Clin Lab Anal. 2021. PMID: 34617636 Free PMC article. Review.
Cited by
-
Circular RNAs in the pathogenesis of SARS-CoV-2: potential diagnostic biomarkers and therapeutic targets.Funct Integr Genomics. 2025 Jan 2;25(1):4. doi: 10.1007/s10142-024-01509-6. Funct Integr Genomics. 2025. PMID: 39745517 Review.
-
Non-coding RNAs expression in SARS-CoV-2 infection: pathogenesis, clinical significance, and therapeutic targets.Signal Transduct Target Ther. 2023 Dec 6;8(1):441. doi: 10.1038/s41392-023-01669-0. Signal Transduct Target Ther. 2023. PMID: 38057315 Free PMC article. Review.
-
Computational identification of differentially-expressed genes as suggested novel COVID-19 biomarkers: A bioinformatics analysis of expression profiles.Comput Struct Biotechnol J. 2023;21:3339-3354. doi: 10.1016/j.csbj.2023.06.007. Epub 2023 Jun 12. Comput Struct Biotechnol J. 2023. PMID: 37347079 Free PMC article.
-
Ferroptosis and circular RNAs: new horizons in cancer therapy.EXCLI J. 2024 Apr 25;23:570-599. doi: 10.17179/excli2024-7005. eCollection 2024. EXCLI J. 2024. PMID: 38887390 Free PMC article. Review.
-
CircRNA: a rising therapeutic strategy for lung injury induced by pulmonary toxicants.Arch Toxicol. 2024 May;98(5):1297-1310. doi: 10.1007/s00204-024-03706-5. Epub 2024 Mar 18. Arch Toxicol. 2024. PMID: 38498160 Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

