Hedgehog Signaling Regulates Treg to Th17 Conversion Through Metabolic Rewiring in Breast Cancer
- PMID: 37058110
- PMCID: PMC10159910
- DOI: 10.1158/2326-6066.CIR-22-0426
Hedgehog Signaling Regulates Treg to Th17 Conversion Through Metabolic Rewiring in Breast Cancer
Abstract
The tumor immune microenvironment dynamically evolves to support tumor growth and progression. Immunosuppressive regulatory T cells (Treg) promote tumor growth and metastatic seeding in patients with breast cancer. Deregulation of plasticity between Treg and Th17 cells creates an immune regulatory framework that enables tumor progression. Here, we discovered a functional role for Hedgehog (Hh) signaling in promoting Treg differentiation and immunosuppressive activity, and when Hh activity was inhibited, Tregs adopted a Th17-like phenotype complemented by an enhanced inflammatory profile. Mechanistically, Hh signaling promoted O-GlcNAc modifications of critical Treg and Th17 transcription factors, Foxp3 and STAT3, respectively, that orchestrated this transition. Blocking Hh reprogramed Tregs metabolically, dampened their immunosuppressive activity, and supported their transdifferentiation into inflammatory Th17 cells that enhanced the recruitment of cytotoxic CD8+ T cells into tumors. Our results demonstrate a previously unknown role for Hh signaling in the regulation of Treg differentiation and activity and the switch between Tregs and Th17 cells in the tumor microenvironment.
©2023 American Association for Cancer Research.
Conflict of interest statement
Figures

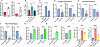
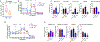
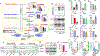

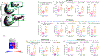
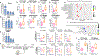
Similar articles
-
Sirtuin2 suppresses the polarization of regulatory T cells toward T helper 17 cells through repressing the expression of signal transducer and activator of transcription 3 in a mouse colitis model.Immun Inflamm Dis. 2024 Feb;12(2):e1160. doi: 10.1002/iid3.1160. Immun Inflamm Dis. 2024. PMID: 38415949 Free PMC article.
-
Suppressive IL-17A+Foxp3+ and ex-Th17 IL-17AnegFoxp3+ Treg cells are a source of tumour-associated Treg cells.Nat Commun. 2017 Mar 14;8:14649. doi: 10.1038/ncomms14649. Nat Commun. 2017. PMID: 28290453 Free PMC article.
-
MondoA-Thioredoxin-Interacting Protein Axis Maintains Regulatory T-Cell Identity and Function in Colorectal Cancer Microenvironment.Gastroenterology. 2021 Aug;161(2):575-591.e16. doi: 10.1053/j.gastro.2021.04.041. Epub 2021 Apr 24. Gastroenterology. 2021. PMID: 33901495
-
The role of T helper 17 and regulatory T cells in tumor microenvironment.Immunopharmacol Immunotoxicol. 2019 Feb;41(1):16-24. doi: 10.1080/08923973.2019.1566925. Epub 2019 Feb 4. Immunopharmacol Immunotoxicol. 2019. PMID: 30714422 Review.
-
Emerging roles of T helper 17 and regulatory T cells in lung cancer progression and metastasis.Mol Cancer. 2016 Oct 27;15(1):67. doi: 10.1186/s12943-016-0551-1. Mol Cancer. 2016. PMID: 27784305 Free PMC article. Review.
Cited by
-
Regulatory T cell-associated gene signature correlates with prognostic risk and immune infiltration in patients with breast cancer.Transl Cancer Res. 2024 Dec 31;13(12):6766-6781. doi: 10.21037/tcr-24-1118. Epub 2024 Dec 27. Transl Cancer Res. 2024. PMID: 39816556 Free PMC article.
-
Ribosomal RNA Biosynthesis Functionally Programs Tumor-Associated Macrophages to Support Breast Cancer Progression.Cancer Res. 2025 Apr 15;85(8):1459-1478. doi: 10.1158/0008-5472.CAN-24-0707. Cancer Res. 2025. PMID: 39903832
-
Hedgehog pathway and cancer: A new area (Review).Oncol Rep. 2024 Sep;52(3):116. doi: 10.3892/or.2024.8775. Epub 2024 Jul 12. Oncol Rep. 2024. PMID: 38994763 Free PMC article. Review.
-
Interactions between hedgehog signaling pathway and the complex tumor microenvironment in breast cancer: current knowledge and therapeutic promises.Cell Commun Signal. 2024 Sep 9;22(1):432. doi: 10.1186/s12964-024-01812-6. Cell Commun Signal. 2024. PMID: 39252010 Free PMC article. Review.
-
Pleiotropic physiological functions of Piezo1 in human body and its effect on malignant behavior of tumors.Front Physiol. 2024 Apr 16;15:1377329. doi: 10.3389/fphys.2024.1377329. eCollection 2024. Front Physiol. 2024. PMID: 38690080 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

