The Alzheimer's disease risk factor INPP5D restricts neuroprotective microglial responses in amyloid beta-mediated pathology
- PMID: 37061460
- PMCID: PMC10576836
- DOI: 10.1002/alz.13089
The Alzheimer's disease risk factor INPP5D restricts neuroprotective microglial responses in amyloid beta-mediated pathology
Abstract
Introduction: Mutations in INPP5D, which encodes for the SH2-domain-containing inositol phosphatase SHIP-1, have recently been linked to an increased risk of developing late-onset Alzheimer's disease. While INPP5D expression is almost exclusively restricted to microglia in the brain, little is known regarding how SHIP-1 affects neurobiology or neurodegenerative disease pathogenesis.
Methods: We generated and investigated 5xFAD Inpp5dfl/fl Cx3cr1Ert2Cre mice to ascertain the function of microglial SHIP-1 signaling in response to amyloid beta (Aβ)-mediated pathology.
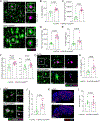
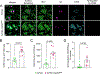
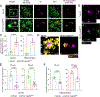
Results: SHIP-1 deletion in microglia led to substantially enhanced recruitment of microglia to Aβ plaques, altered microglial gene expression, and marked improvements in neuronal health. Further, SHIP-1 loss enhanced microglial plaque containment and Aβ engulfment when compared to microglia from Cre-negative 5xFAD Inpp5dfl/fl littermate controls.
Discussion: These results define SHIP-1 as a pivotal regulator of microglial responses during Aβ-driven neurological disease and suggest that targeting SHIP-1 may offer a promising strategy to treat Alzheimer's disease.
Highlights: Inpp5d deficiency in microglia increases plaque-associated microglia numbers. Loss of Inpp5d induces activation and phagocytosis transcriptional pathways. Plaque encapsulation and engulfment by microglia are enhanced with Inpp5d deletion. Genetic ablation of Inpp5d protects against plaque-induced neuronal dystrophy.
Keywords: Alzheimer's disease; INPP5D; SHIP-1; amyloid beta; amyloidosis; disease-associated microglia; microglia; neurodegenerative disease; neuroimmunology.
© 2023 The Authors. Alzheimer's & Dementia published by Wiley Periodicals LLC on behalf of Alzheimer's Association.
Conflict of interest statement
Figures


References
-
- Alzheimer’s Association. More Than Normal Aging: Understanding Mild Cognitive Impairment. Facts and Figures 2022.
-
- Keren-Shaul H, Spinrad A, Weiner A, Matcovitch-Natan O, Dvir-Szternfeld R, Ulland TK, et al. A Unique Microglia Type Associated with Restricting Development of Alzheimer’s Disease. Cell 2017;169:1276–1290.e17. 10.1016/J.CELL.2017.05.018/ATTACHMENT/63D76284-8C53-4D00-93AA-E1E70B1AE6DE/MMC7.XLSX. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

