The first Brazilian bovine breed: structure and genetic diversity of the Curraleiro Pé-duro
- PMID: 37065705
- PMCID: PMC10103694
- DOI: 10.7717/peerj.14768
The first Brazilian bovine breed: structure and genetic diversity of the Curraleiro Pé-duro
Abstract
Background: The production of animal-based foods from native breeds have a synergistic relationship with the regional culture, the local climate, and mainly the maintenance of alternative genetic resources for a system with a lower environmental impact. Thus the efficiency of conservation and production depends on assessing the variability of these local breeds. In the case of Curraleiro Pé-duro cattle, the most adapted individuals have undergone natural selection over five hundred years in the Brazilian savannas, mating with little or no human interference. The peculiarities of these biomes, where the regional flora is the food base and cattle is raised in extensive areas, likely influenced the genetic composition of the different groups that make up the first cattle breed of Brazil.
Methods: To evaluate the composition, diversity, variation, differentiation, and genetic structure of the populations studied, samples of hair follicles from 474 individuals of different animal categories (calves, yearlings, heifers, cows, and bulls) from three farms, defined as subpopulations "A", "B", and "C", were collected. The animals were genotyped for 17 microsatellite markers using a DNA sequencer. After verification of monomorphic alleles, alleles outside the expected size range, and for the presence of stutter bands, the results were subjected to statistical analysis.
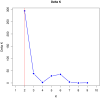
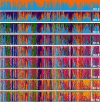
Results: The markers used were suitable for the proposed application with a mean Polymorphism Information Content (PIC) of 0.62. On average, the effective alleles were 4.25 per marker, with mean heterozygosities of 0.74 (observed and expected), which was lower in herd A (0.70) in comparison to herds B (0.77) and C (0.74). The analysis of molecular variance (AMOVA) revealed a higher rate of variation within herds (98.5%) and lower among herds (1.5%) (FSTranging from 0.00723 and 0.03198; p-values < 0.05). However no significant differences among herds where found with the Mantel test based on geographic distances. The formation of genetic clusters of all animals sampled with the software Structure resulted in minimum cluster values, with two main genetic groups (K = 2) observed among the evaluated animals. Therefore, based on PIC and heterozygosity values, a wide genetic diversity was observed, despite little differences in population structure (AMOVA, FST, and Structure results) among sampling sites.
Keywords: AMOVA; Microsatellites; Native breed; Slatikin’s genetic distance.
©2023 Rocha-Silva et al.
Conflict of interest statement
The authors declare there are no competing interests.
Figures
References
-
- Barendse W, Armitage SM, Kossarek LM, Shalom A, Kirkpatrick BW, Ryan AM, Clayton D, Li L, Neibergs HL, Zhang N, Grosse WM, Weiss J, Creighton P, McCarthy F, Ron M, Teale AJ, Fries R, McGraw RA, Moore SS, Georges M, Soller M, Womack JE, Hetzel DJS. A genetic linkage map of the bovine genome. Nature Genetics. 1994;6(3):227–235. doi: 10.1038/ng0394-227. - DOI - PubMed
-
- Cao J, Baumung R, Boettcher P, Scherf B, Besbes B, Leroy G. Monitoring and progress in the implementation of the global plan of action on animal genetic resources. Sustainability. 2021;13(2):775. doi: 10.3390/su13020775. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous