Core-Genome Multilocus Sequence Typing for Epidemiological and Evolutionary Analyses of Phytopathogenic Xanthomonas citri
- PMID: 37067413
- PMCID: PMC10231234
- DOI: 10.1128/aem.02101-22
Core-Genome Multilocus Sequence Typing for Epidemiological and Evolutionary Analyses of Phytopathogenic Xanthomonas citri
Abstract
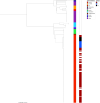
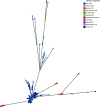
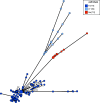
Xanthomonas citri subsp. citri is the cause of bacterial citrus canker, responsible for major economic losses to the citrus industry. X. citri subspecies and pathovars are responsible for diseases in soybean, common bean, mango, pomegranate, and cashew. X. citri disease has been tracked using several typing methods, but recent studies using genomic sequencing have been key to understanding the evolutionary relationships within the species, including fundamental differences among X. citri subsp. citri pathotypes. Here, we describe a core-genome multilocus sequence typing (cgMLST) scheme for X. citri based on 250 genomes comprising multiple examples of X. citri subsp. citri pathotypes A, A*, and Aw; X. citri subsp. malvacearum; X. citri pv. aurantifolii, pv. fuscans, pv. glycines, pv. mangiferaeindicae, pv. viticola, and pv. vignicola; and single isolates of X. citri pv. dieffenbachiae and pv. punicae. This data set included genomic sequencing of 100 novel X. citri subsp. citri isolates. cgMLST, based on 1,618 core genes across 250 genomes, is implemented at PubMLST (https://pubmlst.org/organisms/xanthomonas-citri/). GrapeTree minimum-spanning tree and Interactive Tree of Life (iTOL) neighbor-joining phylogenies generated from the cgMLST data resolved almost identical groupings of isolates to a core-genome single nucleotide polymorphism (SNP)-based neighbor-joining phylogeny. These resolved identical groupings of X. citri subsp. citri pathotypes and X. citri subspecies and pathovars. X. citri cgMLST should prove to be an increasingly valuable resource for the study of this key species of plant-pathogenic bacteria. Users can submit genomic data and associated metadata for comparison with previously characterized isolates at PubMLST to allow the rapid characterization of the local, national, and global epidemiology of these pathogens and examine evolutionary relationships. IMPORTANCE Xanthomonas citri is a plant pathogen that causes major economic losses to the citrus industry and sweet orange production in particular. Several subspecies and pathogens are recognized, with host ranges including soybean, common bean, mango, pomegranate, and cashew, among others. Recent genomic studies have shown that host-adapted X. citri subspecies and pathovars and X. citri subsp. citri pathotypes form distinct clades. In this study, we describe a core-genome multilocus sequence typing (cgMLST) scheme for this species that can rapidly and robustly discriminate among these ecologically distinct, host-adapted clades. We have established this scheme and associated databases containing genomic sequences and metadata at PubMLST, which users can interrogate with their own genome sequences to determine X. citri subspecies, pathovars, and pathotypes. X. citri cgMLST should prove to be an invaluable tool for the study of the epidemiology and evolution of this major plant pathogen.
Keywords: MLST; Xanthomonas citri; cgMLST; citrus canker.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Behlau F, Gochez AM, Jones JB. 2020. Diversity and copper resistance of Xanthomonas affecting citrus. Trop Plant Pathol 45:200–212. doi: 10.1007/s40858-020-00340-1. - DOI
-
- Ference CM, Gochez AM, Behlau F, Wang N, Graham JH, Jones JB. 2018. Recent advances in the understanding of Xanthomonas citri ssp. citri pathogenesis and citrus canker disease management: Xcc pathogenesis and citrus canker management. Mol Plant Pathol 19:1302–1318. doi: 10.1111/mpp.12638. - DOI - PMC - PubMed
-
- Gordon JL, Lefeuvre P, Escalon A, Barbe V, Cruveiller S, Gagnevin L, Pruvost O. 2015. Comparative genomics of 43 strains of Xanthomonas citri pv. citri reveals the evolutionary events giving rise to pathotypes with different host ranges. BMC Genomics 16:1098. doi: 10.1186/s12864-015-2310-x. - DOI - PMC - PubMed
Publication types
MeSH terms
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources

