Maize β-amylase7 encodes 2 proteins using alternative transcriptional start sites: Nuclear BAM7 and plastidic BAM2
- PMID: 37067898
- PMCID: PMC10400034
- DOI: 10.1093/plphys/kiad227
Maize β-amylase7 encodes 2 proteins using alternative transcriptional start sites: Nuclear BAM7 and plastidic BAM2
Abstract
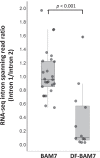
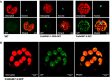
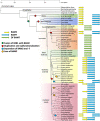
An unusual β-amylase7 (BAM7) gene in some angiosperms, including grasses such as maize (Zea mays), appears to encode 2 functionally distinct proteins: a nuclear-localized transcription factor (BAM7) and a plastid-localized starch hydrolase (BAM2). In Arabidopsis (Arabidopsis thaliana), these 2 proteins are encoded by separate genes on different chromosomes but their physiological functions are not well established. Using the maize BAM7 gene as a model, we detected 2 populations of transcripts by 5'-RACE which encode the predicted proteins. The 2 transcripts are apparently synthesized independently using separate core promoters about 1 kb apart, the second of which is located in the first intron of the full-length gene. The N-terminus of the shorter protein, ZmBAM7-S, begins near the 3' end of the first intron of ZmBAM7-L and starts with a predicted chloroplast transit peptide. We previously showed that ZmBAM7-S is catalytically active with properties like those of AtBAM2. Here, we report that ZmBAM7-S targets green fluorescent protein to plastids. The transcript encoding the longer protein, ZmBAM7-L, encodes an additional DNA-binding domain containing a functional nuclear localization signal. This putative dual-function gene originated at least 400 Mya, prior to the emergence of ferns, and has persisted in some angiosperms that lack a separate BAM2 gene. It appears to have been duplicated and subfunctionalized in at least 4 lineages of land plants, resulting in 2 genes resembling Arabidopsis BAM2 and BAM7. Targeting of 2 products from a single gene to different subcellular locations is not uncommon in plants, but it is unusual when they are predicted to serve completely different functions in the 2 locations.
© The Author(s) 2023. Published by Oxford University Press on behalf of American Society of Plant Biologists.
Conflict of interest statement
Conflict of interest statement. The authors declare no conflict of interest.
Figures


Comment in
-
Lost in transition: Maize BAM7 is a dual function gene encoding a nuclear BAM7 and plastidial BAM2.Plant Physiol. 2023 Aug 3;192(4):2577-2579. doi: 10.1093/plphys/kiad260. Plant Physiol. 2023. PMID: 37139856 Free PMC article. No abstract available.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

