Digital Histopathology by Infrared Spectroscopic Imaging
- PMID: 37068745
- PMCID: PMC10408309
- DOI: 10.1146/annurev-anchem-101422-090956
Digital Histopathology by Infrared Spectroscopic Imaging
Abstract
Infrared (IR) spectroscopic imaging records spatially resolved molecular vibrational spectra, enabling a comprehensive measurement of the chemical makeup and heterogeneity of biological tissues. Combining this novel contrast mechanism in microscopy with the use of artificial intelligence can transform the practice of histopathology, which currently relies largely on human examination of morphologic patterns within stained tissue. First, this review summarizes IR imaging instrumentation especially suited to histopathology, analyses of its performance, and major trends. Second, an overview of data processing methods and application of machine learning is given, with an emphasis on the emerging use of deep learning. Third, a discussion on workflows in pathology is provided, with four categories proposed based on the complexity of methods and the analytical performance needed. Last, a set of guidelines, termed experimental and analytical specifications for spectroscopic imaging in histopathology, are proposed to help standardize the diversity of approaches in this emerging area.
Keywords: chemical imaging; deep learning; digital pathology; imaging; infrared spectroscopy; machine learning.
Figures



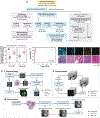
References
-
- Kumar V, Abbas AK, Aster JC. 2015. Robbins and Cotran Pathologic Basis of Disease. Philadelphia: Elsevier/Saunders. 9th ed.
-
- Wood BR, Quinn MA, Tait B, Ashdown M, Hislop T, et al. 1998. FTIR microspectroscopic study of cell types and potential confounding variables in screening for cervical malignancies. Biospectroscopy 4:75–91 - PubMed
-
- Blout ER, Mellors RC. 1949. Infrared spectra of tissues. Science 110:137–38 - PubMed
-
- Jackson M, Sowa MG, Mantsch HH. 1997. Infrared spectroscopy: a new frontier in medicine. Biophys. Chem 68:109–25 - PubMed
-
- Diem M, Boydston-White S, Chiriboga L. 1999. Infrared spectroscopy of cells and tissues: shining light onto a novel subject. Appl. Spectrosc 53:148A–61A
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

