Proteomics: Progress and Promise of High-Throughput Proteomics in Chronic Kidney Disease
- PMID: 37076045
- PMCID: PMC10326701
- DOI: 10.1016/j.mcpro.2023.100550
Proteomics: Progress and Promise of High-Throughput Proteomics in Chronic Kidney Disease
Abstract
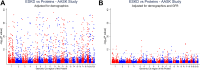
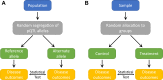
Current proteomic tools permit the high-throughput analysis of the blood proteome in large cohorts, including those enriched for chronic kidney disease (CKD) or its risk factors. To date, these studies have identified numerous proteins associated with cross-sectional measures of kidney function, as well as with the longitudinal risk of CKD progression. Representative signals that have emerged from the literature include an association between levels of testican-2 and favorable kidney prognosis and an association between levels of TNFRSF1A and TNFRSF1B and worse kidney prognosis. For these and other associations, however, understanding whether the proteins play a causal role in kidney disease pathogenesis remains a fundamental challenge, especially given the strong impact that kidney function can have on blood protein levels. Prior to investing in dedicated animal models or randomized trials, methods that leverage the availability of genotyping in epidemiologic cohorts-including Mendelian randomization, colocalization analyses, and proteome-wide association studies-can add evidence for causal inference in CKD proteomics research. In addition, integration of large-scale blood proteome analyses with urine and tissue proteomics, as well as improved assessment of posttranslational protein modifications (e.g., carbamylation), represent important future directions. Taken together, these approaches seek to translate progress in large-scale proteomic profiling into the promise of improved diagnostic tools and therapeutic target identification in kidney disease.
Keywords: Mendelian randomization; Proteomics; causal inference; chronic kidney disease; high-throughput; pQTL.
Copyright © 2023 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare no competing interests.
Figures


References
-
- Gramolini A., Lau E., Liu P.P. Identifying low-abundance biomarkers: aptamer-based proteomics potentially enables more sensitive detection in cardiovascular diseases. Circulation. 2016;134:286–289. - PubMed
-
- Ferkingstad E., Sulem P., Atlason B.A., Sveinbjornsson G., Magnusson M.I., Styrmisdottir E.L., et al. Large-scale integration of the plasma proteome with genetics and disease. Nat. Genet. 2021;53:1712–1721. - PubMed
-
- Sun B.B., Chiou J., Traylor M., Benner C., Hsu Y.-H., Richardson T.G., et al. Genetic regulation of the human plasma proteome in 54,306 UK biobank participants. bioRxiv. 2022 doi: 10.1101/2022.06.17.496443. - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

