Plasma metabolomic characterization of SARS-CoV-2 Omicron infection
- PMID: 37076483
- PMCID: PMC10113737
- DOI: 10.1038/s41419-023-05791-3
Plasma metabolomic characterization of SARS-CoV-2 Omicron infection
Abstract
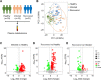
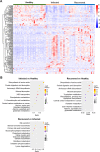
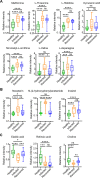
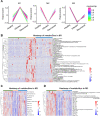
Omicron variants of SARS-CoV-2 have spread rapidly worldwide; however, most infected patients have mild or no symptoms. This study aimed to understand the host response to Omicron infections by performing metabolomic profiling of plasma. We observed that Omicron infections triggered an inflammatory response and innate immune, and adaptive immunity was suppressed, including reduced T-cell response and immunoglobulin antibody production. Similar to the original SARS-CoV-2 strain circulating in 2019, the host developed an anti-inflammatory response and accelerated energy metabolism in response to Omicron infection. However, differential regulation of macrophage polarization and reduced neutrophil function has been observed in Omicron infections. Interferon-induced antiviral immunity was not as strong in Omicron infections as in the original SARS-CoV-2 infections. The host response to Omicron infections increased antioxidant capacity and liver detoxification more than in the original strain. Hence, these findings suggest that Omicron infections cause weaker inflammatory alterations and immune responses than the original SARS-CoV-2 strain.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
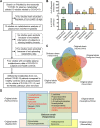
References
-
- Fall A, Eldesouki RE, Sachithanandham J, Morris CP, Norton JM, Gaston DC, et al. The displacement of the SARS-CoV-2 variant Delta with Omicron: an investigation of hospital admissions and upper respiratory viral loads. EBioMedicine. 2022;79:104008. doi: 10.1016/j.ebiom.2022.104008. - DOI - PMC - PubMed
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

