CTCF is a DNA-tension-dependent barrier to cohesin-mediated loop extrusion
- PMID: 37076620
- PMCID: PMC10132984
- DOI: 10.1038/s41586-023-05961-5
CTCF is a DNA-tension-dependent barrier to cohesin-mediated loop extrusion
Abstract
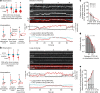
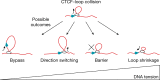
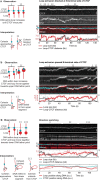
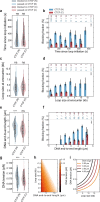
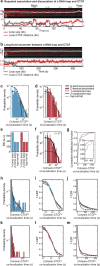
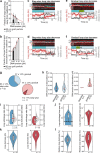
In eukaryotes, genomic DNA is extruded into loops by cohesin1. By restraining this process, the DNA-binding protein CCCTC-binding factor (CTCF) generates topologically associating domains (TADs)2,3 that have important roles in gene regulation and recombination during development and disease1,4-7. How CTCF establishes TAD boundaries and to what extent these are permeable to cohesin is unclear8. Here, to address these questions, we visualize interactions of single CTCF and cohesin molecules on DNA in vitro. We show that CTCF is sufficient to block diffusing cohesin, possibly reflecting how cohesive cohesin accumulates at TAD boundaries, and is also sufficient to block loop-extruding cohesin, reflecting how CTCF establishes TAD boundaries. CTCF functions asymmetrically, as predicted; however, CTCF is dependent on DNA tension. Moreover, CTCF regulates cohesin's loop-extrusion activity by changing its direction and by inducing loop shrinkage. Our data indicate that CTCF is not, as previously assumed, simply a barrier to cohesin-mediated loop extrusion but is an active regulator of this process, whereby the permeability of TAD boundaries can be modulated by DNA tension. These results reveal mechanistic principles of how CTCF controls loop extrusion and genome architecture.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures








Comment in
-
CTCF works better under tension.Nat Rev Mol Cell Biol. 2023 Jun;24(6):379. doi: 10.1038/s41580-023-00614-x. Nat Rev Mol Cell Biol. 2023. PMID: 37161005 No abstract available.
Similar articles
-
Permeable TAD boundaries and their impact on genome-associated functions.Bioessays. 2024 Oct;46(10):e2400137. doi: 10.1002/bies.202400137. Epub 2024 Aug 2. Bioessays. 2024. PMID: 39093600
-
Cohesin distribution alone predicts chromatin organization in yeast via conserved-current loop extrusion.Genome Biol. 2024 Nov 14;25(1):293. doi: 10.1186/s13059-024-03432-2. Genome Biol. 2024. PMID: 39543681 Free PMC article.
-
High-resolution CTCF footprinting reveals impact of chromatin state on cohesin extrusion.Nat Commun. 2025 May 15;16(1):4506. doi: 10.1038/s41467-025-57775-w. Nat Commun. 2025. PMID: 40374602 Free PMC article.
-
CTCF as a boundary factor for cohesin-mediated loop extrusion: evidence for a multi-step mechanism.Nucleus. 2020 Dec;11(1):132-148. doi: 10.1080/19491034.2020.1782024. Nucleus. 2020. PMID: 32631111 Free PMC article. Review.
-
Pushing the TAD boundary: Decoding insulator codes of clustered CTCF sites in 3D genomes.Bioessays. 2024 Oct;46(10):e2400121. doi: 10.1002/bies.202400121. Epub 2024 Aug 21. Bioessays. 2024. PMID: 39169755 Review.
Cited by
-
CTCF deletion alters the pluripotency and DNA methylation profile of human iPSCs.Front Cell Dev Biol. 2023 Nov 30;11:1302448. doi: 10.3389/fcell.2023.1302448. eCollection 2023. Front Cell Dev Biol. 2023. PMID: 38099298 Free PMC article.
-
Thermodynamic principles link in vitro transcription factor affinities to single-molecule chromatin states in cells.bioRxiv [Preprint]. 2025 Jan 30:2025.01.27.635162. doi: 10.1101/2025.01.27.635162. bioRxiv. 2025. PMID: 39975040 Free PMC article. Preprint.
-
Evolution and function of chromatin domains across the tree of life.Nat Struct Mol Biol. 2024 Dec;31(12):1824-1837. doi: 10.1038/s41594-024-01427-y. Epub 2024 Nov 26. Nat Struct Mol Biol. 2024. PMID: 39592879 Review.
-
Emerging roles of cohesin-STAG2 in cancer.Oncogene. 2025 Feb;44(5):277-287. doi: 10.1038/s41388-024-03221-y. Epub 2024 Nov 29. Oncogene. 2025. PMID: 39613934 Review.
-
Atypical Modes of CTCF Binding Facilitate Tissue-Specific and Neuronal Activity-Dependent Gene Expression States.Mol Neurobiol. 2024 Jun;61(6):3240-3257. doi: 10.1007/s12035-023-03762-5. Epub 2023 Nov 18. Mol Neurobiol. 2024. PMID: 37979036
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

