Ovulatory signal-triggered chromatin remodeling in ovarian granulosa cells by HDAC2 phosphorylation activation-mediated histone deacetylation
- PMID: 37076890
- PMCID: PMC10116676
- DOI: 10.1186/s13072-023-00485-8
Ovulatory signal-triggered chromatin remodeling in ovarian granulosa cells by HDAC2 phosphorylation activation-mediated histone deacetylation
Abstract
Background: Epigenetic reprogramming is involved in luteinizing hormone (LH)-induced ovulation; however, the underlying mechanisms are largely unknown.
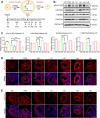
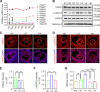
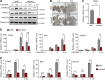
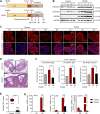
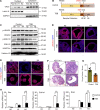
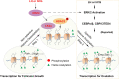
Results: We here observed a rapid histone deacetylation process between two waves of active transcription mediated by the follicle-stimulating hormone (FSH) and the LH congener human chorionic gonadotropin (hCG), respectively. Analysis of the genome-wide H3K27Ac distribution in hCG-treated granulosa cells revealed that a rapid wave of genome-wide histone deacetylation remodels the chromatin, followed by the establishment of specific histone acetylation for ovulation. HDAC2 phosphorylation activation coincides with histone deacetylation in mouse preovulatory follicles. When HDAC2 was silenced or inhibited, histone acetylation was retained, leading to reduced gene transcription, retarded cumulus expansion, and ovulation defect. HDAC2 phosphorylation was associated with CK2α nuclear translocation, and inhibition of CK2α attenuated HDAC2 phosphorylation, retarded H3K27 deacetylation, and inactivated the ERK1/2 signaling cascade.
Conclusions: This study demonstrates that the ovulatory signal erases histone acetylation through activation of CK2α-mediated HDAC2 phosphorylation in granulosa cells, which is an essential prerequisite for subsequent successful ovulation.
Keywords: Chromatin remodeling; Granulosa cells; H3K27Ac; HDAC2; Luteinizing hormone; Ovulation.
© 2023. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

References
-
- Richards JS, Jahnsen T, Hedin L, Lifka J, Ratoosh S, Durica JM, Goldring NB. Ovarian follicular development: from physiology to molecular biology. Recent Prog Horm Res. 1987;43:231–276. - PubMed
-
- Park JY, Su YQ, Ariga M, Law E, Jin SL, Conti M: EGF-like growth factors as mediators of LH action in the ovulatory follicle. Science (80- ) 2004, 303(5658):682–684. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

