Severe acute respiratory syndrome coronavirus-2 accessory proteins ORF3a and ORF7a modulate autophagic flux and Ca2+ homeostasis in yeast
- PMID: 37077240
- PMCID: PMC10106705
- DOI: 10.3389/fmicb.2023.1152249
Severe acute respiratory syndrome coronavirus-2 accessory proteins ORF3a and ORF7a modulate autophagic flux and Ca2+ homeostasis in yeast
Abstract
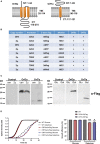
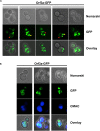
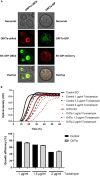
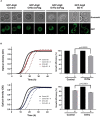
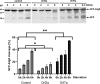
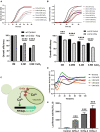
Virus infection involves the manipulation of key host cell functions by specialized virulence proteins. The Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) small accessory proteins ORF3a and ORF7a have been implicated in favoring virus replication and spreading by inhibiting the autophagic flux within the host cell. Here, we apply yeast models to gain insights into the physiological functions of both SARS-CoV-2 small open reading frames (ORFs). ORF3a and ORF7a can be stably overexpressed in yeast cells, producing a decrease in cellular fitness. Both proteins show a distinguishable intracellular localization. ORF3a localizes to the vacuolar membrane, whereas ORF7a targets the endoplasmic reticulum. Overexpression of ORF3a and ORF7a leads to the accumulation of Atg8 specific autophagosomes. However, the underlying mechanism is different for each viral protein as assessed by the quantification of the autophagic degradation of Atg8-GFP fusion proteins, which is inhibited by ORF3a and stimulated by ORF7a. Overexpression of both SARS-CoV-2 ORFs decreases cellular fitness upon starvation conditions, where autophagic processes become essential. These data confirm previous findings on SARS-CoV-2 ORF3a and ORF7a manipulating autophagic flux in mammalian cell models and are in agreement with a model where both small ORFs have synergistic functions in stimulating intracellular autophagosome accumulation, ORF3a by inhibiting autophagosome processing at the vacuole and ORF7a by promoting autophagosome formation at the ER. ORF3a has an additional function in Ca2+ homeostasis. The overexpression of ORF3a confers calcineurin-dependent Ca2+ tolerance and activates a Ca2+ sensitive FKS2-luciferase reporter, suggesting a possible ORF3a-mediated Ca2+ efflux from the vacuole. Taken together, we show that viral accessory proteins can be functionally investigated in yeast cells and that SARS-CoV-2 ORF3a and ORF7a proteins interfere with autophagosome formation and processing as well as with Ca2+ homeostasis from distinct cellular targets.
Keywords: Ca2+ homeostasis; ORF3a; ORF7a; SARS-CoV-2; Saccharomyces cerevisiae; autophagy.
Copyright © 2023 Garrido-Huarte, Fita-Torró, Viana, Pascual-Ahuir and Proft.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Manipulation of autophagy by SARS-CoV-2 proteins.Autophagy. 2021 Sep;17(9):2659-2661. doi: 10.1080/15548627.2021.1953847. Epub 2021 Jul 19. Autophagy. 2021. PMID: 34281462 Free PMC article.
-
The ORF7a protein of SARS-CoV-2 initiates autophagy and limits autophagosome-lysosome fusion via degradation of SNAP29 to promote virus replication.Autophagy. 2023 Feb;19(2):551-569. doi: 10.1080/15548627.2022.2084686. Epub 2022 Jun 19. Autophagy. 2023. PMID: 35670302 Free PMC article.
-
SARS-CoV-2 ORF7a blocked autophagy flux by intervening in the fusion between autophagosome and lysosome to promote viral infection and pathogenesis.J Med Virol. 2023 Nov;95(11):e29200. doi: 10.1002/jmv.29200. J Med Virol. 2023. PMID: 37916857
-
Regulation of autophagy by SARS-CoV-2: The multifunctional contributions of ORF3a.J Med Virol. 2023 Jul;95(7):e28959. doi: 10.1002/jmv.28959. J Med Virol. 2023. PMID: 37485696 Review.
-
The role of SARS-CoV-2 accessory proteins in immune evasion.Biomed Pharmacother. 2022 Dec;156:113889. doi: 10.1016/j.biopha.2022.113889. Epub 2022 Oct 17. Biomed Pharmacother. 2022. PMID: 36265309 Free PMC article. Review.
Cited by
-
Is Autophagy a Friend or Foe in SARS-CoV-2 Infection?Viruses. 2024 Sep 20;16(9):1491. doi: 10.3390/v16091491. Viruses. 2024. PMID: 39339967 Free PMC article. Review.
-
Backstage Heroes-Yeast in COVID-19 Research.Int J Mol Sci. 2024 Nov 25;25(23):12661. doi: 10.3390/ijms252312661. Int J Mol Sci. 2024. PMID: 39684373 Free PMC article. Review.
-
Mechanism of Lactiplantibacillus plantarum regulating Ca2+ affecting the replication of PEDV in small intestinal epithelial cells.Front Microbiol. 2023 Sep 26;14:1251275. doi: 10.3389/fmicb.2023.1251275. eCollection 2023. Front Microbiol. 2023. PMID: 37840713 Free PMC article.
-
SARS-CoV-2 ORF3a drives dynamic dense body formation for optimal viral infectivity.Nat Commun. 2025 May 12;16(1):4393. doi: 10.1038/s41467-025-59475-x. Nat Commun. 2025. PMID: 40355429 Free PMC article.
-
SARS-CoV-2 ORF3a drives dynamic dense body formation for optimal viral infectivity.Res Sq [Preprint]. 2024 May 17:rs.3.rs-4292014. doi: 10.21203/rs.3.rs-4292014/v1. Res Sq. 2024. Update in: Nat Commun. 2025 May 12;16(1):4393. doi: 10.1038/s41467-025-59475-x. PMID: 38798602 Free PMC article. Updated. Preprint.
References
LinkOut - more resources
Full Text Sources
Miscellaneous

