Relevance of the Trillion-Sized Chemical Space "eXplore" as a Source for Drug Discovery
- PMID: 37077402
- PMCID: PMC10108389
- DOI: 10.1021/acsmedchemlett.3c00021
Relevance of the Trillion-Sized Chemical Space "eXplore" as a Source for Drug Discovery
Abstract
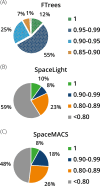
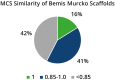
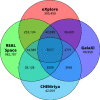
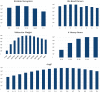
Within the past two decades, virtual combinatorial compound collections, so-called chemical spaces, became an important molecule source for pharmaceutical research all over the world. The emergence of compound vendor chemical spaces with rapidly growing numbers of molecules raises questions about their application suitability and the quality of the content. Here, we examine the composition of the recently published and, so far, biggest chemical space, "eXplore", which comprises approximately 2.8 trillion virtual product molecules. The utility of eXplore to retrieve interesting chemistry around approved drugs and common Bemis Murcko scaffolds has been assessed with several methods (FTrees, SpaceLight, SpaceMACS). Further, the overlap between several vendor chemical spaces and a physicochemical property distribution analysis has been performed. Despite the straightforward chemical reactions underlying its setup, eXplore is demonstrated to provide relevant and, most importantly, easily accessible molecules for drug discovery campaigns.
© 2023 American Chemical Society.
Conflict of interest statement
The authors declare the following competing financial interest(s): A.N. and R.K. are employees of BioSolveIT, a company offering paid software to screen the eXplore space. L.M. is an employee of eMolecules that offers building blocks and screening compounds described in this work.
Figures


Similar articles
-
Maximum Common Substructure Searching in Combinatorial Make-on-Demand Compound Spaces.J Chem Inf Model. 2022 May 9;62(9):2133-2150. doi: 10.1021/acs.jcim.1c00640. Epub 2021 Sep 3. J Chem Inf Model. 2022. PMID: 34478299
-
Overlap of On-demand Ultra-large Combinatorial Spaces with On-the-shelf Drug-like Libraries.Mol Inform. 2023 Jan;42(1):e2200163. doi: 10.1002/minf.202200163. Epub 2022 Sep 30. Mol Inform. 2023. PMID: 36072995
-
Exploiting cheminformatic and machine learning to navigate the available chemical space of potential small molecule inhibitors of SARS-CoV-2.Comput Struct Biotechnol J. 2021;19:424-438. doi: 10.1016/j.csbj.2020.12.028. Epub 2020 Dec 29. Comput Struct Biotechnol J. 2021. PMID: 33391634 Free PMC article.
-
Charting, navigating, and populating natural product chemical space for drug discovery.J Med Chem. 2012 Jul 12;55(13):5989-6001. doi: 10.1021/jm300288g. Epub 2012 May 11. J Med Chem. 2012. PMID: 22537178 Review.
-
Expanding the medicinally relevant chemical space with compound libraries.Drug Discov Today. 2012 Jul;17(13-14):718-26. doi: 10.1016/j.drudis.2012.04.001. Epub 2012 Apr 10. Drug Discov Today. 2012. PMID: 22515962 Review.
Cited by
-
AI is a viable alternative to high throughput screening: a 318-target study.Sci Rep. 2024 Apr 2;14(1):7526. doi: 10.1038/s41598-024-54655-z. Sci Rep. 2024. PMID: 38565852 Free PMC article.
-
Protein Structure-Based Organic Chemistry-Driven Ligand Design from Ultralarge Chemical Spaces.ACS Cent Sci. 2024 Feb 13;10(3):615-627. doi: 10.1021/acscentsci.3c01521. eCollection 2024 Mar 27. ACS Cent Sci. 2024. PMID: 38559302 Free PMC article.
-
Chemical space as a unifying theme for chemistry.J Cheminform. 2025 Jan 16;17(1):6. doi: 10.1186/s13321-025-00954-0. J Cheminform. 2025. PMID: 39825400 Free PMC article.
-
CoLiNN: A Tool for Fast Chemical Space Visualization of Combinatorial Libraries Without Enumeration.Mol Inform. 2025 Mar;44(3):e202400263. doi: 10.1002/minf.202400263. Mol Inform. 2025. PMID: 40099935 Free PMC article.
-
Synthesis of Complex Tetracyclic Fused Scaffolds Enabled by (3 + 2) Cycloaddition.Org Lett. 2024 Jun 14;26(23):4873-4876. doi: 10.1021/acs.orglett.4c01269. Epub 2024 May 31. Org Lett. 2024. PMID: 38820198 Free PMC article.
References
-
- Sadybekov A. A.; Sadybekov A. V.; Liu Y.; Iliopoulos-Tsoutsouvas C.; Huang X.-P.; Pickett J.; Houser B.; Patel N.; Tran N. K.; Tong F.; Zvonok N.; Jain M. K.; Savych O.; Radchenko D. S.; Nikas S. P.; Petasis N. A.; Moroz Y. S.; Roth B. L.; Makriyannis A.; Katritch V. Synthon-Based Ligand Discovery in Virtual Libraries of over 11 Billion Compounds. Nature 2022, 601, 452–459. 10.1038/s41586-021-04220-9. - DOI - PMC - PubMed
-
- Muller J.; Klein R.; Tarkhanova O.; Gryniukova A.; Borysko P.; Merkl S.; Ruf M.; Neumann A.; Gastreich M.; Moroz Y. S.; Klebe G.; Glinca S. Magnet for the Needle in Haystacks: “Crystal Structure First” Fragment Hits Unlock Active Chemical Matter Using Targeted Exploration of Vast Chemical Spaces. J. Med. Chem. 2022, 65 (23), 15663–15678. 10.1021/acs.jmedchem.2c00813. - DOI - PubMed
-
- Beroza P.; Crawford J. J.; Ganichkin O.; Gendelev L.; Harris S. F.; Klein R.; Miu A.; Steinbacher S.; Klingler F.-M.; Lemmen C. Chemical Space Docking Finds Novel ROCK1 Kinase Inhibitors by Large-Scale Structure-Based Virtual Screening. Nat. Commun. 2022, 6447.10.1038/s41467-022-33981-8. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

