Distinct subpopulations of DN1 thymocytes exhibit preferential γδ T lineage potential
- PMID: 37077921
- PMCID: PMC10106834
- DOI: 10.3389/fimmu.2023.1106652
Distinct subpopulations of DN1 thymocytes exhibit preferential γδ T lineage potential
Abstract
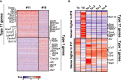
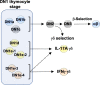
The αβ and γδ T cell lineages both differentiate in the thymus from common uncommitted progenitors. The earliest stage of T cell development is known as CD4-CD8- double negative 1 (DN1), which has previously been shown to be a heterogenous mixture of cells. Of these, only the CD117+ fraction has been proposed to be true T cell progenitors that progress to the DN2 and DN3 thymocyte stages, at which point the development of the αβ and γδ T cell lineages diverge. However, recently, it has been shown that at least some γδ T cells may be derived from a subset of CD117- DN thymocytes. Along with other ambiguities, this suggests that T cell development may not be as straightforward as previously thought. To better understand early T cell development, particularly the heterogeneity of DN1 thymocytes, we performed a single cell RNA sequence (scRNAseq) of mouse DN and γδ thymocytes and show that the various DN stages indeed comprise a transcriptionally diverse subpopulations of cells. We also show that multiple subpopulations of DN1 thymocytes exhibit preferential development towards the γδ lineage. Furthermore, specific γδ-primed DN1 subpopulations preferentially develop into IL-17 or IFNγ-producing γδ T cells. We show that DN1 subpopulations that only give rise to IL-17-producing γδ T cells already express many of the transcription factors associated with type 17 immune cell responses, while the DN1 subpopulations that can give rise to IFNγ-producing γδ T cell already express transcription factors associated with type 1 immune cell responses.
Keywords: T cell development; gamma delta (γδ) T cells; lineage decision; scRNAseq; thymocyte.
Copyright © 2023 Oh, Liu, Tomei, Luo, Skinner, Berzins, Naik, Gray and Chong.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures






References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

