CD103+ regulatory T cells underlie resistance to radio-immunotherapy and impair CD8+ T cell activation in glioblastoma
- PMID: 37081259
- PMCID: PMC10212765
- DOI: 10.1038/s43018-023-00547-6
CD103+ regulatory T cells underlie resistance to radio-immunotherapy and impair CD8+ T cell activation in glioblastoma
Abstract
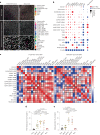
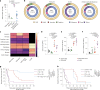
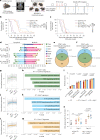
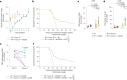
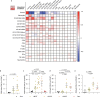
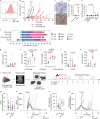
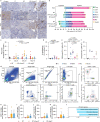
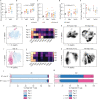
Glioblastomas are aggressive primary brain tumors with an inherent resistance to T cell-centric immunotherapy due to their low mutational burden and immunosuppressive tumor microenvironment. Here we report that fractionated radiotherapy of preclinical glioblastoma models induce a tenfold increase in T cell content. Orthogonally, spatial imaging mass cytometry shows T cell enrichment in human recurrent tumors compared with matched primary glioblastoma. In glioblastoma-bearing mice, α-PD-1 treatment applied at the peak of T cell infiltration post-radiotherapy results in a modest survival benefit compared with concurrent α-PD-1 administration. Following α-PD-1 therapy, CD103+ regulatory T cells (Tregs) with upregulated lipid metabolism accumulate in the tumor microenvironment, and restrain immune checkpoint blockade response by repressing CD8+ T cell activation. Treg targeting elicits tertiary lymphoid structure formation, enhances CD4+ and CD8+ T cell frequency and function and unleashes radio-immunotherapeutic efficacy. These results support the rational design of therapeutic regimens limiting the induction of immunosuppressive feedback pathways in the context of T cell immunotherapy in glioblastoma.
© 2023. The Author(s).
Conflict of interest statement
The authors have no competing interests to declare.
Figures





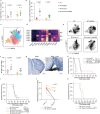
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

