Autophagy-prominent cell clusters among human lens epithelial cells: integrated single-cell RNA-sequencing analysis
- PMID: 37081480
- PMCID: PMC10116761
- DOI: 10.1186/s12886-023-02910-8
Autophagy-prominent cell clusters among human lens epithelial cells: integrated single-cell RNA-sequencing analysis
Abstract
Background: Autophagy is an important process that maintains the quality of intracellular proteins and organelles. There is extensive evidence that autophagy has an important role in the lens. Human lens epithelial cells (HLECs) play a key role in the internal homeostasis of the lens. HLEC subtypes have been identified, but autophagy-prominent cell clusters among HLECs have not been characterized.
Purpose: To explore the existence of autophagy-prominent cell clusters in HLECs.
Methods: Three donated lenses (HLECs from two whole lenses and HLECs from one lens without the anterior central 6-mm zone) were used for single-cell RNA sequencing (scRNA-seq). AUCell and AddModuleScore analysis were used to identify potential autophagy-prominent cell clusters. Transmission electron microscopy (TEM) was used to confirm the results.
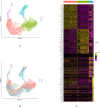
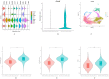
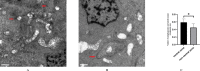
Results: High-quality transcripts from 18,120 cells were acquired by scRNA-seq of the two intact lenses. Unsupervised clustering classified the cells into four clusters. AUCell and AddModuleScore analysis revealed cluster 1 is autophagy-prominent. scRNA-seq analysis of HLECs from the lens capsule lacking the central zone confirmed the cluster 1 HLECs was located in the central capsule zone. The TEM result showed that greater autophagy activity was observed in the HLECs in central capsule zone, which further supported the above conclusions based on scRNA-seq analysis that autophagy was prominent in the central zone where the cluster 1 HLECs located.
Conclusions: We identified an autophagy-prominent cell cluster among HLECs and revealed that it was localized in the central zone of the lens capsule. Our findings will aid investigations of autophagy in HLECs and provide insights to guide related research.
Keywords: Autophagy; Human lens epithelial cells; Single-cell RNA-sequencing; Transmission electron microscopy.
© 2023. The Author(s).
Conflict of interest statement
The authors have no conflicts of interest to declare.
Figures

Similar articles
-
Presence of a transcriptionally active glucocorticoid receptor alpha in lens epithelial cells.Invest Ophthalmol Vis Sci. 2003 Dec;44(12):5269-76. doi: 10.1167/iovs.03-0401. Invest Ophthalmol Vis Sci. 2003. PMID: 14638726
-
Capsule-epithelium-fibre unit ultrastructure in the human lens.Int J Med Sci. 2023 Feb 27;20(4):542-550. doi: 10.7150/ijms.82446. eCollection 2023. Int J Med Sci. 2023. PMID: 37057214 Free PMC article.
-
Metformin attenuates the epithelial-mesenchymal transition of lens epithelial cells through the AMPK/TGF-β/Smad2/3 signalling pathway.Exp Eye Res. 2021 Nov;212:108763. doi: 10.1016/j.exer.2021.108763. Epub 2021 Sep 10. Exp Eye Res. 2021. PMID: 34517004
-
Effects of senescent lens epithelial cells on the severity of age-related cortical cataract in humans: A case-control study.Medicine (Baltimore). 2016 Jun;95(25):e3869. doi: 10.1097/MD.0000000000003869. Medicine (Baltimore). 2016. PMID: 27336873 Free PMC article.
-
Impacts of autophagy on the formation of organelle-free zone during the lens development.Mol Biol Rep. 2023 May;50(5):4551-4564. doi: 10.1007/s11033-023-08323-6. Epub 2023 Mar 6. Mol Biol Rep. 2023. PMID: 36877352 Review.
Cited by
-
miR-24-3p mediates Keap1/Nrf2 axis to promote autophagy and thereby inhibit lens epithelial cell early senescence.Int Ophthalmol. 2025 Aug 9;45(1):327. doi: 10.1007/s10792-025-03701-4. Int Ophthalmol. 2025. PMID: 40782127
-
Integrated analysis of single-cell RNA-seq and ATAC-seq in lens epithelial cells: Unveiling the role of ATF6 as a key transcription factor.Genes Dis. 2025 Mar 22;12(6):101610. doi: 10.1016/j.gendis.2025.101610. eCollection 2025 Nov. Genes Dis. 2025. PMID: 40837408 Free PMC article.
-
Integrated single-cell multiomics uncovers foundational regulatory mechanisms of lens development and pathology.Development. 2024 Jan 1;151(1):dev202249. doi: 10.1242/dev.202249. Epub 2024 Jan 5. Development. 2024. PMID: 38180241 Free PMC article.
-
Single-Cell RNA Sequencing Analysis of the Early Postnatal Mouse Lens Epithelium.Invest Ophthalmol Vis Sci. 2023 Oct 3;64(13):37. doi: 10.1167/iovs.64.13.37. Invest Ophthalmol Vis Sci. 2023. PMID: 37870847 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

