Phenotypic Characterization and Antibiograms of Extended-Spectrum Beta-Lactamase-Producing Escherichia coli Isolated at the Human-Animal-Environment Interface Using a One Health Approach Among Households in Wakiso District, Uganda
- PMID: 37081947
- PMCID: PMC10112474
- DOI: 10.2147/IDR.S398951
Phenotypic Characterization and Antibiograms of Extended-Spectrum Beta-Lactamase-Producing Escherichia coli Isolated at the Human-Animal-Environment Interface Using a One Health Approach Among Households in Wakiso District, Uganda
Abstract
Background: The occurrence of extended spectrum beta-lactamase (ESBL) producing bacteria such as Escherichia coli has increasingly become recognized beyond hospital settings. Resistance to other types of antibiotics limits treatment options while the existence of such bacteria among humans, animals, and the environment is suggestive of potential zoonotic and reverse-zoonotic transmission. This study aimed to establish the antibiotic susceptibility profiles of the ESBL-producing Escherichia coli (ESBL-EC) from human, animal, and environmental isolates obtained among farming households within Wakiso district using a One Health approach.
Methods: A total of 100 ESBL-EC isolates from humans 35/100 (35%), animals 56/100 (56%), and the environment 9/100 (9%) were tested for susceptibility to 11 antibiotics. This was done using the Kirby-Bauer disk diffusion method according to Clinical and Laboratory Standards Institute (CLSI) guidelines. Data were analyzed in STATA ver. 16 and graphs were drawn in Microsoft excel ver. 10.
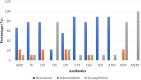
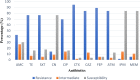
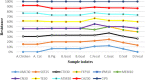
Results: Most of the ESBL-EC isolates (98%) were resistant to more than two antibiotics. ESBL-EC isolates were most susceptible to meropenem (MEM) (88.0%), and imipenem (82.0%) followed by gentamicin (72%). ESBL-EC isolates from humans were most susceptible to meropenem (MEM) followed by imipenem (IPM)> gentamicin (CN)> ciprofloxacin (CIP). Animal samples were more susceptible to MEM, IPM, and CN but were highly resistant to cefotaxime (CTX)> cefepime (FEP)>other antibiotics. Multidrug resistance (MDR) was mostly reported among households keeping goats under intensive husbandry practices. Seven percent of the isolates exhibited carbapenem resistance while 22% showed aminoglycoside resistance. Similar resistance patterns among humans, animals, and environmental samples were also reported.
Conclusion: Our study provides baseline information on non-hospital-based MDR caused by ESBL-EC using a One Health approach. ESBL-EC isolates were prevalent among apparently healthy community members, animals, and their environment. It is important to conduct more One Health approach studies to generate evidence on the drivers, resistance patterns, and transmission of ESBL-producing organisms at the human-animal-environmental interface.
Keywords: ESBL-producing; Escherichia coli; Wakiso; antibiogram; community; one health.
© 2023 Muleme et al.
Conflict of interest statement
The authors report no conflicts of interest in this work.
Figures



Similar articles
-
[Investigation of beta-lactamase genes and clonal relationship among the extended-spectrum beta-lactamase producing nosocomial Escherichia coli isolates].Mikrobiyol Bul. 2015 Jan;49(1):15-25. doi: 10.5578/mb.8437. Mikrobiyol Bul. 2015. PMID: 25706727 Turkish.
-
Molecular characterization and antibiotic resistance profile of ESBL-producing Escherichia coli isolated from healthy cow raw milk in smallholder dairy farms in Bangladesh.Vet World. 2023 Jun;16(6):1333-1339. doi: 10.14202/vetworld.2023.1333-1339. Epub 2023 Jun 13. Vet World. 2023. PMID: 37577207 Free PMC article.
-
[Investigation of the susceptibilities of extended-spectrum beta-lactamase-producing Escherichia coli and Klebsiella spp. strains to ertapenem and other carbapenems].Mikrobiyol Bul. 2011 Jan;45(1):28-35. Mikrobiyol Bul. 2011. PMID: 21341156 Turkish.
-
Extended-Spectrum ß-Lactamase-Producing Escherichia coli Among Humans, Beef Cattle, and Abattoir Environments in Nigeria.Front Cell Infect Microbiol. 2022 Apr 7;12:869314. doi: 10.3389/fcimb.2022.869314. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 35463650 Free PMC article.
-
Measures used to assess the burden of ESBL-producing Escherichia coli infections in humans: a scoping review.JAC Antimicrob Resist. 2021 Feb 14;3(1):dlaa104. doi: 10.1093/jacamr/dlaa104. eCollection 2021 Mar. JAC Antimicrob Resist. 2021. PMID: 34223063 Free PMC article.
Cited by
-
The Global Rise of ESBL-Producing Escherichia coli in the Livestock Sector: A Five-Year Overview.Animals (Basel). 2024 Aug 27;14(17):2490. doi: 10.3390/ani14172490. Animals (Basel). 2024. PMID: 39272275 Free PMC article. Review.
-
Carbapenem Resistance in Animal-Environment-Food from Africa: A Systematic Review, Recommendations and Perspectives.Infect Drug Resist. 2024 May 3;17:1699-1728. doi: 10.2147/IDR.S458317. eCollection 2024. Infect Drug Resist. 2024. PMID: 38715963 Free PMC article. Review.
References
-
- Cassini A, Högberg LD, Plachouras D, et al. Attributable deaths and disability-adjusted life-years caused by infections with antibiotic-resistant bacteria in the EU and the European Economic Area in 2015: a population-level modelling analysis. Lancet Infect Dis. 2019;19:56–66. doi:10.1016/S1473-3099(18)30605-4 - DOI - PMC - PubMed
-
- World Health Organization. WHO Report on Surveillance of Antibiotic Consumption: 2016–2018 Early Implementation. Geneva: World Health Organization; 2018.
LinkOut - more resources
Full Text Sources

