Integrated analysis of SR-like protein kinases Sky1 and Sky2 links signaling networks with transcriptional regulation in Candida albicans
- PMID: 37082713
- PMCID: PMC10111165
- DOI: 10.3389/fcimb.2023.1108235
Integrated analysis of SR-like protein kinases Sky1 and Sky2 links signaling networks with transcriptional regulation in Candida albicans
Abstract
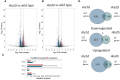
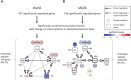
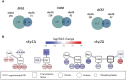
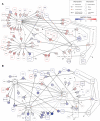
Fungal infections are a major global health burden where Candida albicans is among the most common fungal pathogen in humans and is a common cause of invasive candidiasis. Fungal phenotypes, such as those related to morphology, proliferation and virulence are mainly driven by gene expression, which is primarily regulated by kinase signaling cascades. Serine-arginine (SR) protein kinases are highly conserved among eukaryotes and are involved in major transcriptional processes in human and S. cerevisiae. Candida albicans harbors two SR protein kinases, while Sky2 is important for metabolic adaptation, Sky1 has similar functions as in S. cerevisiae. To investigate the role of these SR kinases for the regulation of transcriptional responses in C. albicans, we performed RNA sequencing of sky1Δ and sky2Δ and integrated a comprehensive phosphoproteome dataset of these mutants. Using a Systems Biology approach, we study transcriptional regulation in the context of kinase signaling networks. Transcriptomic enrichment analysis indicates that pathways involved in the regulation of gene expression are downregulated and mitochondrial processes are upregulated in sky1Δ. In sky2Δ, primarily metabolic processes are affected, especially for arginine, and we observed that arginine-induced hyphae formation is impaired in sky2Δ. In addition, our analysis identifies several transcription factors as potential drivers of the transcriptional response. Among these, a core set is shared between both kinase knockouts, but it appears to regulate different subsets of target genes. To elucidate these diverse regulatory patterns, we created network modules by integrating the data of site-specific protein phosphorylation and gene expression with kinase-substrate predictions and protein-protein interactions. These integrated signaling modules reveal shared parts but also highlight specific patterns characteristic for each kinase. Interestingly, the modules contain many proteins involved in fungal morphogenesis and stress response. Accordingly, experimental phenotyping shows a higher resistance to Hygromycin B for sky1Δ. Thus, our study demonstrates that a combination of computational approaches with integration of experimental data can offer a new systems biological perspective on the complex network of signaling and transcription. With that, the investigation of the interface between signaling and transcriptional regulation in C. albicans provides a deeper insight into how cellular mechanisms can shape the phenotype.
Keywords: Candida albicans; kinase signaling; network analysis; phosphoproteome; sky kinases; transcriptional regulation; transcriptome.
Copyright © 2023 Luther, Brandt, Vylkova, Dandekar, Müller and Dittrich.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Candida albicans SR-Like Protein Kinases Regulate Different Cellular Processes: Sky1 Is Involved in Control of Ion Homeostasis, While Sky2 Is Important for Dipeptide Utilization.Front Cell Infect Microbiol. 2022 May 6;12:850531. doi: 10.3389/fcimb.2022.850531. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 35601106 Free PMC article.
-
The Ndr/LATS Kinase Cbk1 Regulates a Specific Subset of Ace2 Functions and Suppresses the Hypha-to-Yeast Transition in Candida albicans.mBio. 2020 Aug 18;11(4):e01900-20. doi: 10.1128/mBio.01900-20. mBio. 2020. PMID: 32817109 Free PMC article.
-
Lipid Signaling via Pkh1/2 Regulates Fungal CO2 Sensing through the Kinase Sch9.mBio. 2017 Jan 31;8(1):e02211-16. doi: 10.1128/mBio.02211-16. mBio. 2017. PMID: 28143980 Free PMC article.
-
Stress-Activated Protein Kinases in Human Fungal Pathogens.Front Cell Infect Microbiol. 2019 Jul 17;9:261. doi: 10.3389/fcimb.2019.00261. eCollection 2019. Front Cell Infect Microbiol. 2019. PMID: 31380304 Free PMC article. Review.
-
Co-regulation of pathogenesis with dimorphism and phenotypic switching in Candida albicans, a commensal and a pathogen.Int J Med Microbiol. 2002 Oct;292(5-6):299-311. doi: 10.1078/1438-4221-00215. Int J Med Microbiol. 2002. PMID: 12452278 Review.
Cited by
-
Generation of a high confidence set of domain-domain interface types to guide protein complex structure predictions by AlphaFold.Bioinformatics. 2024 Aug 2;40(8):btae482. doi: 10.1093/bioinformatics/btae482. Bioinformatics. 2024. PMID: 39171834 Free PMC article.
-
The Antidepressant Sertraline Modulates Gene Expression and Alternative Splicing Events in the Dermatophyte Trichophyton rubrum: A Comprehensive Analysis.Genes (Basel). 2025 Jan 24;16(2):146. doi: 10.3390/genes16020146. Genes (Basel). 2025. PMID: 40004476 Free PMC article.
-
Histone deacetylase Sir2 promotes the systemic Candida albicans infection by facilitating its immune escape via remodeling the cell wall and maintaining the metabolic activity.mBio. 2024 Jun 12;15(6):e0044524. doi: 10.1128/mbio.00445-24. Epub 2024 Apr 29. mBio. 2024. PMID: 38682948 Free PMC article.
-
Associations between Genomic Variants and Antifungal Susceptibilities in the Archived Global Candida auris Population.J Fungi (Basel). 2024 Jan 22;10(1):86. doi: 10.3390/jof10010086. J Fungi (Basel). 2024. PMID: 38276031 Free PMC article.
-
Systematic analysis of the Candida albicans kinome reveals environmentally contingent protein kinase-mediated regulation of filamentation and biofilm formation in vitro and in vivo.mBio. 2024 Aug 14;15(8):e0124924. doi: 10.1128/mbio.01249-24. Epub 2024 Jul 1. mBio. 2024. PMID: 38949302 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

