Methylthioadenosine phosphorylase deficiency in tumors: A compelling therapeutic target
- PMID: 37091983
- PMCID: PMC10113547
- DOI: 10.3389/fcell.2023.1173356
Methylthioadenosine phosphorylase deficiency in tumors: A compelling therapeutic target
Abstract
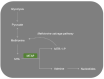
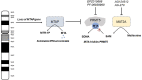
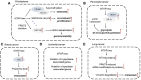
The methionine salvage pathway is responsible for recycling sulfur-containing metabolites to methionine. This salvage pathway has been found to be implicated in cell apoptosis, proliferation, differentiation and inflammatory response. Methylthioadenosine phosphorylase (MTAP) catalyzes the reversible phosphorolysis of 5'-methylthioadenosine, a by-product produced from polyamine biosynthesis. The MTAP gene is located adjacent to the cyclin-dependent kinase inhibitor 2A gene and co-deletes with CDKN2A in nearly 15% of tumors. Moreover, MTAP-deleted tumor cells exhibit greater sensitivity to methionine depletion and to the inhibitors of purine synthesis. In this review, we first summarized the molecular structure and expression of MTAP in tumors. Furthermore, we discussed PRMT5 and MAT2A as a potential vulnerability for MTAP-deleted tumors. The complex and dynamic role of MTAP in diverse malignancies has also been discussed. Finally, we demonstrated the implications for the treatment of MTAP-deleted tumors.
Keywords: CDKN2A; MTAP; glioma; methionine salvage pathway; tumor.
Copyright © 2023 Fan, Zhang and Zou.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Disordered methionine metabolism in MTAP/CDKN2A-deleted cancers leads to dependence on PRMT5.Science. 2016 Mar 11;351(6278):1208-13. doi: 10.1126/science.aad5944. Epub 2016 Feb 11. Science. 2016. PMID: 26912361
-
MAT2A Inhibition Blocks the Growth of MTAP-Deleted Cancer Cells by Reducing PRMT5-Dependent mRNA Splicing and Inducing DNA Damage.Cancer Cell. 2021 Feb 8;39(2):209-224.e11. doi: 10.1016/j.ccell.2020.12.010. Epub 2021 Jan 14. Cancer Cell. 2021. PMID: 33450196
-
MTAP Deletions in Cancer Create Vulnerability to Targeting of the MAT2A/PRMT5/RIOK1 Axis.Cell Rep. 2016 Apr 19;15(3):574-587. doi: 10.1016/j.celrep.2016.03.043. Epub 2016 Apr 7. Cell Rep. 2016. PMID: 27068473
-
Targeting tumors that lack methylthioadenosine phosphorylase (MTAP) activity: current strategies.Cancer Biol Ther. 2011 Apr 1;11(7):627-32. doi: 10.4161/cbt.11.7.14948. Epub 2011 Apr 1. Cancer Biol Ther. 2011. PMID: 21301207 Free PMC article. Review.
-
The potential and challenges of targeting MTAP-negative cancers beyond synthetic lethality.Front Oncol. 2023 Sep 19;13:1264785. doi: 10.3389/fonc.2023.1264785. eCollection 2023. Front Oncol. 2023. PMID: 37795443 Free PMC article. Review.
Cited by
-
Landscape of targets within nucleoside metabolism for the modification of immune responses.Front Oncol. 2025 May 30;15:1483769. doi: 10.3389/fonc.2025.1483769. eCollection 2025. Front Oncol. 2025. PMID: 40519286 Free PMC article. Review.
-
5'-S-(3-Aminophenyl)-5'-thioadenosine, a Novel Chemoprotective Agent for Reducing Toxic Side Effects of Fluorouracil in Treatment of MTAP-Deficient Cancers.Mol Cancer Ther. 2025 Jul 2;24(7):1030-1039. doi: 10.1158/1535-7163.MCT-24-0656. Mol Cancer Ther. 2025. PMID: 40062378
-
Next-generation therapies for pancreatic cancer.Expert Rev Gastroenterol Hepatol. 2024 Jan-Feb;18(1-3):55-72. doi: 10.1080/17474124.2024.2322648. Epub 2024 Feb 28. Expert Rev Gastroenterol Hepatol. 2024. PMID: 38415709 Free PMC article. Review.
-
Targeting amino acid in tumor therapy.Front Oncol. 2025 Jun 4;15:1582116. doi: 10.3389/fonc.2025.1582116. eCollection 2025. Front Oncol. 2025. PMID: 40535116 Free PMC article. Review.
-
Chemically programmed metabolism drives a superior cell fitness for cartilage regeneration.Sci Adv. 2024 Sep 13;10(37):eadp4408. doi: 10.1126/sciadv.adp4408. Epub 2024 Sep 11. Sci Adv. 2024. PMID: 39259800 Free PMC article.
References
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

