Adaptation of Prokaryotic Toxins for Negative Selection and Cloning-Independent Markerless Mutagenesis in Streptococcus Species
- PMID: 37093065
- PMCID: PMC10286703
- DOI: 10.1128/msphere.00682-22
Adaptation of Prokaryotic Toxins for Negative Selection and Cloning-Independent Markerless Mutagenesis in Streptococcus Species
Abstract
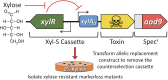
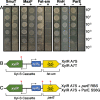
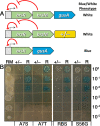
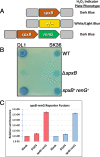
The Streptococcus mutans genetic system offers a variety of strategies to rapidly engineer targeted chromosomal mutations. Previously, we reported the first S. mutans negative selection system that functions in a wild-type background. This system utilizes induced sensitivity to the toxic amino acid analog p-chlorophenylalanine (4-CP) as a negative selection mechanism and was developed for counterselection-based cloning-independent markerless mutagenesis (CIMM). While we have employed this system extensively for our ongoing genetic studies, we have encountered a couple limitations with the system, mainly its narrow host range and the requirement for selection on a toxic substrate. Here, we report the development of a new negative selection system that addresses both limitations, while still retaining the utility of the previous 4-CP-based markerless mutagenesis system. We placed a variety of toxin-encoding genes under the control of the xylose-inducible gene expression cassette (Xyl-S) and found the Fst-sm and ParE toxins to be suitable candidates for inducible negative selection. We combined the inducible toxins with an antibiotic resistance gene to create several different counterselection cassettes. The most broadly useful of these contained a wild-type fst-sm open reading frame transcriptionally fused to a point mutant form of the Xyl-S expression system, which we subsequently named IFDC4. IFDC4 was shown to exhibit exceptionally low background resistance, with 3- to 4-log reductions in cell number observed when plating on xylose-supplemented medium. IFDC4 also functioned similarly in multiple strains of S. mutans as well as with Streptococcus gordonii and Streptococcus sanguinis. We performed CIMM with IFDC4 and successfully engineered a variety of different types of markerless mutations in all three species. The counterselection strategy described here provides a template approach that should be adaptable for the creation of similar counterselection systems in many other bacteria. IMPORTANCE Multiple medically significant Streptococcus species, such as S. mutans, have highly sophisticated genetic systems available, largely as a consequence of their amenability to genetic manipulation via natural competence. Despite this, few options are available for the creation of markerless mutations in streptococci, especially within wild-type strains. Markerless mutagenesis is a critical tool for genetic studies, as it allows the user to explore many fundamental questions that are not easily addressable using marked mutagenesis. Here, we describe a new approach for streptococcal markerless mutagenesis that offers a variety of advantages over the current approach, which employs induced sensitivity to the toxic substrate 4-CP. The approach employed here should be readily adaptable for the creation of similar markerless mutagenesis systems in other organisms.
Keywords: Streptococcus; counterselection; markerless mutation; negative selection; toxin-antitoxin; xylose.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
