Identifying TME signatures for cervical cancer prognosis based on GEO and TCGA databases
- PMID: 37095983
- PMCID: PMC10121839
- DOI: 10.1016/j.heliyon.2023.e15096
Identifying TME signatures for cervical cancer prognosis based on GEO and TCGA databases
Abstract
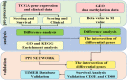
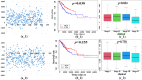
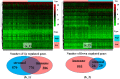
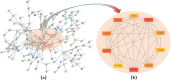
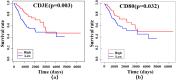
The mortality rate from cervical cancer (CESC), a malignant tumor that affects women, has increased significantly globally in recent years. The discovery of biomarkers points to a direction for the diagnosis of cervical cancer with the advancement of bioinformatics technology. The goal of this study was to look for potential biomarkers for the diagnosis and prognosis of CESC using the GEO and TCGA databases. Because of the high dimension and small sample size of the omic data, or the use of biomarkers generated from a single omic data, the diagnosis of cervical cancer may be inaccurate and unreliable. The purpose of this study was to search the GEO and TCGA databases for potential biomarkers for the diagnosis and prognosis of CESC. We begin by downloading CESC (GSE30760) DNA methylation data from GEO, then perform differential analysis on the downloaded methylation data and screen out the differential genes. Then, using estimation algorithms, we score immune cells and stromal cells in the tumor microenvironment and perform survival analysis on the gene expression profile data and the most recent clinical data of CESC from TCGA. Then, using the 'limma' package and Venn plot in R language to perform differential analysis of genes and screen out overlapping genes, these overlapping genes were then subjected to GO and KEGG functional enrichment analysis. The differential genes screened by the GEO methylation data and the differential genes screened by the TCGA gene expression data were intersected to screen out the common differential genes. A protein-protein interaction (PPI) network of gene expression data was then created in order to discover important genes. The PPI network's key genes were crossed with previously identified common differential genes to further validate them. The Kaplan-Meier curve was then used to determine the prognostic importance of the key genes. Survival analysis has shown that CD3E and CD80 are important for the identification of cervical cancer and can be considered as potential biomarkers for cervical cancer.
Keywords: Biomarkers; Cervical cancer; DNA methylation; Differentially expressed genes; Tumor microenvironment.
©2023PublishedbyElsevierLtd.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures




Similar articles
-
Identification of immune related molecular subtypes and prognosis model for predicting prognosis, drug resistance in cervical squamous cell carcinoma.Front Genet. 2023 Mar 17;14:1137995. doi: 10.3389/fgene.2023.1137995. eCollection 2023. Front Genet. 2023. PMID: 37007965 Free PMC article.
-
Methylation-driven genes and their prognostic value in cervical squamous cell carcinoma.Ann Transl Med. 2020 Jul;8(14):868. doi: 10.21037/atm-19-4577. Ann Transl Med. 2020. PMID: 32793712 Free PMC article.
-
Prognostic value of tumour microenvironment-related genes by TCGA database in rectal cancer.J Cell Mol Med. 2021 Jun;25(12):5811-5822. doi: 10.1111/jcmm.16547. Epub 2021 May 5. J Cell Mol Med. 2021. PMID: 33949771 Free PMC article.
-
Ferroptosis-related genes identify tumor immune microenvironment characterization for the prediction of prognosis in cervical cancer.Ann Transl Med. 2022 Jan;10(2):123. doi: 10.21037/atm-21-6265. Ann Transl Med. 2022. PMID: 35282071 Free PMC article.
-
Tumor microenvironment characterization in cervical cancer identifies prognostic relevant gene signatures.PLoS One. 2021 Apr 26;16(4):e0249374. doi: 10.1371/journal.pone.0249374. eCollection 2021. PLoS One. 2021. PMID: 33901225 Free PMC article.
Cited by
-
Comprehensive Single-Cell RNA Sequencing Analysis of Cervical Cancer: Insights Into Tumor Microenvironment and Gene Expression Dynamics.Int J Genomics. 2025 May 13;2025:5027347. doi: 10.1155/ijog/5027347. eCollection 2025. Int J Genomics. 2025. PMID: 40395456 Free PMC article.
-
Mendelian randomization reveals immune cell composition as a key determinant of cervical cancer prognosis.Discov Oncol. 2025 Apr 29;16(1):635. doi: 10.1007/s12672-025-02455-w. Discov Oncol. 2025. PMID: 40299140 Free PMC article.
-
Integrative Analysis of Neutrophil-Associated Genes Reveals Prognostic Significance and Immune Microenvironment Modulation in Cervical Cancer.Biomedicines. 2025 May 30;13(6):1348. doi: 10.3390/biomedicines13061348. Biomedicines. 2025. PMID: 40564066 Free PMC article.
-
UPP1 is a dual biomarker of prognosis and immune microenvironment in IDH wild-type glioblastoma.Sci Rep. 2025 Aug 28;15(1):31677. doi: 10.1038/s41598-025-16907-4. Sci Rep. 2025. PMID: 40877486 Free PMC article.
-
Methods in DNA methylation array dataset analysis: A review.Comput Struct Biotechnol J. 2024 May 17;23:2304-2325. doi: 10.1016/j.csbj.2024.05.015. eCollection 2024 Dec. Comput Struct Biotechnol J. 2024. PMID: 38845821 Free PMC article. Review.
References
-
- Small W., Jr., Bacon M.A., Bajaj A., Chuang L.T., Fisher B.J., Harkenrider M.M., Jhingran A., Kitchener H.C., Mileshkin L.R., Viswanathan A.N., Gaffney D.K. Cervical cancer: a global health crisis. Cancer. 2017;123:2404–2412. - PubMed
-
- Twardella D., Geiss K., Radespiel-Troger M., Benner A., Ficker J.H., Meyer M. [Trends in incidence of lung cancer according to histological subtype among men and women in Germany : analysis of cancer registry data with the application of multiple imputation techniques] Bundesgesundheitsblatt - Gesundheitsforsch. - Gesundheitsschutz. 2018;61:20–31. - PubMed
-
- Di J., Rutherford S., Chu C. Review of the cervical cancer burden and population-based cervical cancer screening in China. Asian Pac. J. Cancer Prev. APJCP. 2015;16:7401–7407. - PubMed
LinkOut - more resources
Full Text Sources

