Engineering lanthipeptides by introducing a large variety of RiPP modifications to obtain new-to-nature bioactive peptides
- PMID: 37096385
- PMCID: PMC10373908
- DOI: 10.1093/femsre/fuad017
Engineering lanthipeptides by introducing a large variety of RiPP modifications to obtain new-to-nature bioactive peptides
Abstract
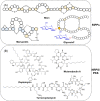
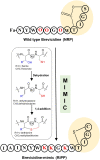
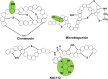
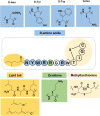
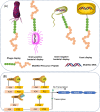
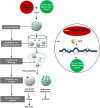
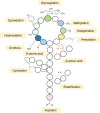
Natural bioactive peptide discovery is a challenging and time-consuming process. However, advances in synthetic biology are providing promising new avenues in peptide engineering that allow for the design and production of a large variety of new-to-nature peptides with enhanced or new bioactivities, using known peptides as templates. Lanthipeptides are ribosomally synthesized and post-translationally modified peptides (RiPPs). The modularity of post-translational modification (PTM) enzymes and ribosomal biosynthesis inherent to lanthipeptides enables their engineering and screening in a high-throughput manner. The field of RiPPs research is rapidly evolving, with many novel PTMs and their associated modification enzymes being identified and characterized. The modularity presented by these diverse and promiscuous modification enzymes has made them promising tools for further in vivo engineering of lanthipeptides, allowing for the diversification of their structures and activities. In this review, we explore the diverse modifications occurring in RiPPs and discuss the potential applications and feasibility of combining various modification enzymes for lanthipeptide engineering. We highlight the prospect of lanthipeptide- and RiPP-engineering to produce and screen novel peptides, including mimics of potent non-ribosomally produced antimicrobial peptides (NRPs) such as daptomycin, vancomycin, and teixobactin, which offer high therapeutic potential.
Keywords: Engineering; NRPs mimics; lanthipeptides; post-translational modification; ripps; synthetic biology.
© The Author(s) 2023. Published by Oxford University Press on behalf of FEMS.
Conflict of interest statement
None declared.
Figures
Similar articles
-
Advancements in the Application of Ribosomally Synthesized and Post-Translationally Modified Peptides (RiPPs).Biomolecules. 2024 Apr 15;14(4):479. doi: 10.3390/biom14040479. Biomolecules. 2024. PMID: 38672495 Free PMC article.
-
Cell-free biosynthesis and engineering of ribosomally synthesized lanthipeptides.Nat Commun. 2024 May 21;15(1):4336. doi: 10.1038/s41467-024-48726-y. Nat Commun. 2024. PMID: 38773100 Free PMC article.
-
Mimicry of a Non-ribosomally Produced Antimicrobial, Brevicidine, by Ribosomal Synthesis and Post-translational Modification.Cell Chem Biol. 2020 Oct 15;27(10):1262-1271.e4. doi: 10.1016/j.chembiol.2020.07.005. Epub 2020 Jul 23. Cell Chem Biol. 2020. PMID: 32707039
-
Modular Use of the Uniquely Small Ring A of Mersacidin Generates the Smallest Ribosomally Produced Lanthipeptide.ACS Synth Biol. 2022 Sep 16;11(9):3078-3087. doi: 10.1021/acssynbio.2c00343. Epub 2022 Sep 5. ACS Synth Biol. 2022. PMID: 36065523 Free PMC article.
-
Linaridin natural products.Nat Prod Rep. 2020 Sep 23;37(9):1152-1163. doi: 10.1039/c9np00074g. Nat Prod Rep. 2020. PMID: 32484193 Review.
Cited by
-
Discovery, Biosynthesis, and Characterization of Rodencin, a Two-Component Lanthipeptide, Harboring d-Amino Acids Introduced by the Unusual Dehydrogenase RodJA.J Nat Prod. 2024 Oct 25;87(10):2344-2354. doi: 10.1021/acs.jnatprod.4c00170. Epub 2024 Sep 20. J Nat Prod. 2024. PMID: 39302883 Free PMC article.
-
Stabilizing Scaffold for Short Peptides Based on Knottins.Curr Cancer Drug Targets. 2024;24(12):1275-1285. doi: 10.2174/0115680096285288240118090050. Curr Cancer Drug Targets. 2024. PMID: 38357956
-
The untapped potential of actinobacterial lanthipeptides as therapeutic agents.Mol Biol Rep. 2023 Dec;50(12):10605-10616. doi: 10.1007/s11033-023-08880-w. Epub 2023 Nov 7. Mol Biol Rep. 2023. PMID: 37934370 Free PMC article. Review.
-
Embleporicin: A Novel Class I Lanthipeptide from the Actinobacteria Embleya sp. NF3.Antibiotics (Basel). 2024 Dec 5;13(12):1179. doi: 10.3390/antibiotics13121179. Antibiotics (Basel). 2024. PMID: 39766569 Free PMC article.
-
Direct Cloning and Heterologous Expression of the Dmxorosin Biosynthetic Gene Cluster from Streptomyces thermolilacinus SPC6, a Halotolerant Actinomycete Isolated from the Desert in China.Int J Mol Sci. 2025 Feb 11;26(4):1492. doi: 10.3390/ijms26041492. Int J Mol Sci. 2025. PMID: 40003958 Free PMC article.
References
-
- Alexandru-Crivac CN, Umeobika C, Leikoski Net al. . Cyclic peptide production using a macrocyclase with enhanced substrate promiscuity and relaxed recognition determinants. Chem Commun. 2017;53:10656–9. - PubMed
-
- Amacher DE. The primary role of hepatic metabolism in idiosyncratic drug-induced liver injury. Exp Opin Drug Metabol Toxicol. 2012;8:335–47. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

