An annotated chromosome-scale reference genome for Eastern black-eared wheatear (Oenanthe melanoleuca)
- PMID: 37097035
- PMCID: PMC10234393
- DOI: 10.1093/g3journal/jkad088
An annotated chromosome-scale reference genome for Eastern black-eared wheatear (Oenanthe melanoleuca)
Abstract
Pervasive convergent evolution and in part high incidences of hybridization distinguish wheatears (songbirds of the genus Oenanthe) as a versatile system to address questions at the forefront of research on the molecular bases of phenotypic and species diversification. To prepare the genomic resources for this venture, we here generated and annotated a chromosome-scale assembly of the Eastern black-eared wheatear (Oenanthe melanoleuca). This species is part of the Oenanthe hispanica complex that is characterized by convergent evolution of plumage coloration and high rates of hybridization. The long-read-based male nuclear genome assembly comprises 1.04 Gb in 32 autosomes, the Z chromosome, and the mitogenome. The assembly is highly contiguous (contig N50, 12.6 Mb; scaffold N50, 70 Mb), with 96% of the genome assembled at the chromosome level and 95.5% benchmarking universal single-copy orthologs (BUSCO) completeness. The nuclear genome was annotated with 18,143 protein-coding genes and 31,333 mRNAs (annotation BUSCO completeness, 98.0%), and about 10% of the genome consists of repetitive DNA. The annotated chromosome-scale reference genome of Eastern black-eared wheatear provides a crucial resource for research into the genomics of adaptation and speciation in an intriguing group of passerines.
Keywords: Oenanthe hispanica complex; Oenanthe melanoleuca; birds; open-habitat chats; repeat content; transcriptome; transposable elements.
© The Author(s) 2023. Published by Oxford University Press on behalf of The Genetics Society of America.
Conflict of interest statement
Conflicts of interest statement The authors declare no conflict of interest.
Figures

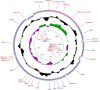

References
-
- Aliabadian M, Kaboli M, Förschler MI, Nijman V, Chamani A, Tillier A, Prodon R, Pasquet E, Ericson PGP, Zuccon D. Convergent evolution of morphological and ecological traits in the open-habitat chat complex (Aves, Muscicapidae: Saxicolinae). Mol Phylogenet Evol. 2012;65(1):35–45. doi: 10.1016/j.ympev.2012.05.011. - DOI - PubMed
-
- Babraham Bioinformatics: Cambridge . FastQC; version 0.10.1: a quality control tool for high throughput sequence data. Cambridge: Babraham Bioinformatics; 2012.
Publication types
MeSH terms
Supplementary concepts
LinkOut - more resources
Full Text Sources
Miscellaneous
