Imaging crossing fibers in mouse, pig, monkey, and human brain using small-angle X-ray scattering
- PMID: 37098400
- PMCID: PMC10811447
- DOI: 10.1016/j.actbio.2023.04.029
Imaging crossing fibers in mouse, pig, monkey, and human brain using small-angle X-ray scattering
Abstract
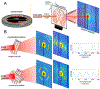
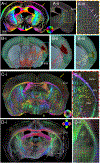
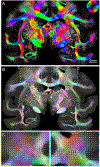
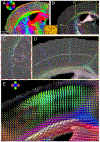
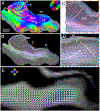
Myelinated axons (nerve fibers) efficiently transmit signals throughout the brain via action potentials. Multiple methods that are sensitive to axon orientations, from microscopy to magnetic resonance imaging, aim to reconstruct the brain's structural connectome. As billions of nerve fibers traverse the brain with various possible geometries at each point, resolving fiber crossings is necessary to generate accurate structural connectivity maps. However, doing so with specificity is a challenging task because signals originating from oriented fibers can be influenced by brain (micro)structures unrelated to myelinated axons. X-ray scattering can specifically probe myelinated axons due to the periodicity of the myelin sheath, which yields distinct peaks in the scattering pattern. Here, we show that small-angle X-ray scattering (SAXS) can be used to detect myelinated, axon-specific fiber crossings. We first demonstrate the capability using strips of human corpus callosum to create artificial double- and triple-crossing fiber geometries, and we then apply the method in mouse, pig, vervet monkey, and human brains. We compare results to polarized light imaging (3D-PLI), tracer experiments, and to outputs from diffusion MRI that sometimes fails to detect crossings. Given its specificity, capability of 3-dimensional sampling and high resolution, SAXS could serve as a ground truth for validating fiber orientations derived using diffusion MRI as well as microscopy-based methods. STATEMENT OF SIGNIFICANCE: To study how the nerve fibers in our brain are interconnected, scientists need to visualize their trajectories, which often cross one another. Here, we show the unique capacity of small-angle X-ray scattering (SAXS) to study these fiber crossings without use of labeling, taking advantage of SAXS's specificity to myelin - the insulating sheath that is wrapped around nerve fibers. We use SAXS to detect double and triple crossing fibers and unveil intricate crossings in mouse, pig, vervet monkey, and human brains. This non-destructive method can uncover complex fiber trajectories and validate other less specific imaging methods (e.g., MRI or microscopy), towards accurate mapping of neuronal connectivity in the animal and human brain.
Keywords: Animal and human brain; Crossing fibers; Diffusion MRI; Fiber orientation mapping; Human hippocampus; Imaging myelinated axons; Mouse/pig/vervet monkey brain; Scanning small-angle X-ray scattering (SAXS).
Copyright © 2023. Published by Elsevier Ltd.
Conflict of interest statement
Declaration of Competing Interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures


Similar articles
-
Using light and X-ray scattering to untangle complex neuronal orientations and validate diffusion MRI.Elife. 2023 May 11;12:e84024. doi: 10.7554/eLife.84024. Elife. 2023. PMID: 37166005 Free PMC article.
-
Scattered Light Imaging: Resolving the substructure of nerve fiber crossings in whole brain sections with micrometer resolution.Neuroimage. 2021 Jun;233:117952. doi: 10.1016/j.neuroimage.2021.117952. Epub 2021 Mar 11. Neuroimage. 2021. PMID: 33716156 Free PMC article.
-
Retrieving neuronal orientations using 3D scanning SAXS and comparison with diffusion MRI.Neuroimage. 2020 Jan 1;204:116214. doi: 10.1016/j.neuroimage.2019.116214. Epub 2019 Sep 27. Neuroimage. 2020. PMID: 31568873 Free PMC article.
-
Small-angle X-ray scattering studies of enzymes.Curr Opin Chem Biol. 2023 Feb;72:102232. doi: 10.1016/j.cbpa.2022.102232. Epub 2022 Nov 30. Curr Opin Chem Biol. 2023. PMID: 36462455 Free PMC article. Review.
-
Advances, Applications, and Perspectives in Small-Angle X-ray Scattering of RNA.Chembiochem. 2023 Sep 1;24(17):e202300110. doi: 10.1002/cbic.202300110. Epub 2023 Jul 19. Chembiochem. 2023. PMID: 37466350 Review.
Cited by
-
Micron-resolution fiber mapping in histology independent of sample preparation.bioRxiv [Preprint]. 2025 Mar 14:2024.03.26.586745. doi: 10.1101/2024.03.26.586745. bioRxiv. 2025. PMID: 38585744 Free PMC article. Preprint.
-
Small-angle scattering tensor tomography algorithm for robust reconstruction of complex textures.Acta Crystallogr A Found Adv. 2023 Nov 1;79(Pt 6):515-526. doi: 10.1107/S205327332300863X. Epub 2023 Oct 19. Acta Crystallogr A Found Adv. 2023. PMID: 37855136 Free PMC article.
-
Using light and X-ray scattering to untangle complex neuronal orientations and validate diffusion MRI.Elife. 2023 May 11;12:e84024. doi: 10.7554/eLife.84024. Elife. 2023. PMID: 37166005 Free PMC article.
References
-
- Morell P, Myelin, 2nd ed., Springer Science+Business Media, New York, 1984, doi:10.1007/978-1-4757-1830-0. - DOI
-
- Hagmann P, From diffusion MRI to brain connectomics, EPFL (2005).
-
- Azevedo FAC, Carvalho LRB, Grinberg LT, Farfel JM, Ferretti REL, Leite REP, Filho WJ, Lent R, Herculano-Houzel S, Equal numbers of neuronal and nonneuronal cells make the human brain an isometrically scaled-up primate brain, J. Comp. Neurol 513 (5) (2009) 532–541, doi:10.1002/cne.21974. - DOI - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

