An aquatic virus exploits the IL6-STAT3-HSP90 signaling axis to promote viral entry
- PMID: 37099596
- PMCID: PMC10166480
- DOI: 10.1371/journal.ppat.1011320
An aquatic virus exploits the IL6-STAT3-HSP90 signaling axis to promote viral entry
Erratum in
-
Correction: An aquatic virus exploits the IL6-STAT3-HSP90 signaling axis to promote viral entry.PLoS Pathog. 2023 Jun 12;19(6):e1011444. doi: 10.1371/journal.ppat.1011444. eCollection 2023 Jun. PLoS Pathog. 2023. PMID: 37307277 Free PMC article.
Abstract
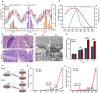
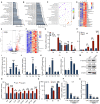
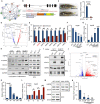
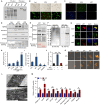
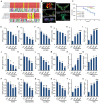
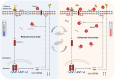
Viral seasonality in the aquaculture industry is an important scientific issue for decades. While the molecular mechanisms underpinning the temperature-dependent pathogenesis of aquatic viral diseases remain largely unknown. Here we report that temperature-dependent activation of IL6-STAT3 signaling was exploited by grass carp reovirus (GCRV) to promote viral entry via increasing the expression of heat shock protein 90 (HSP90). Deploying GCRV infection as a model system, we discovered that GCRV induces the IL6-STAT3-HSP90 signaling activation to achieve temperature-dependent viral entry. Further biochemical and microscopic analyses revealed that the major capsid protein VP7 of GCRV interacted with HSP90 and relevant membrane-associated proteins to boost viral entry. Accordingly, exogenous expression of either IL6, HSP90, or VP7 in cells increased GCRV entry in a dose-dependent manner. Interestingly, other viruses (e.g., koi herpesvirus, Rhabdovirus carpio, Chinese giant salamander iridovirus) infecting ectothermic vertebrates have evolved a similar mechanism to promote their infection. This work delineates a molecular mechanism by which an aquatic viral pathogen exploits the host temperature-related immune response to promote its entry and replication, instructing us on new ways to develop targeted preventives and therapeutics for aquaculture viral diseases.
Copyright: © 2023 Hou et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

