An unexpected hydratase synthesizes the green light-absorbing pigment fucoxanthin
- PMID: 37100425
- PMCID: PMC10396388
- DOI: 10.1093/plcell/koad116
An unexpected hydratase synthesizes the green light-absorbing pigment fucoxanthin
Abstract
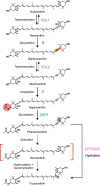
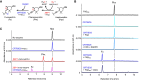
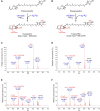
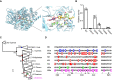
The ketocarotenoid fucoxanthin and its derivatives can absorb blue-green light enriched in marine environments. Fucoxanthin is widely adopted by phytoplankton species as a main light-harvesting pigment, in contrast to land plants that primarily employ chlorophylls. Despite its supreme abundance in the oceans, the last steps of fucoxanthin biosynthesis have remained elusive. Here, we identified the carotenoid isomerase-like protein CRTISO5 as the diatom fucoxanthin synthase that is related to the carotenoid cis-trans isomerase CRTISO from land plants but harbors unexpected enzymatic activity. A crtiso5 knockout mutant in the model diatom Phaeodactylum tricornutum completely lacked fucoxanthin and accumulated the acetylenic carotenoid phaneroxanthin. Recombinant CRTISO5 converted phaneroxanthin into fucoxanthin in vitro by hydrating its carbon-carbon triple bond, instead of functioning as an isomerase. Molecular docking and mutational analyses revealed residues essential for this activity. Furthermore, a photophysiological characterization of the crtiso5 mutant revealed a major structural and functional role of fucoxanthin in photosynthetic pigment-protein complexes of diatoms. As CRTISO5 hydrates an internal alkyne physiologically, the enzyme has unique potential for biocatalytic applications. The discovery of CRTISO5 illustrates how neofunctionalization leads to major diversification events in evolution of photosynthetic mechanisms and the prominent brown coloration of most marine photosynthetic eukaryotes.
© American Society of Plant Biologists 2023. All rights reserved. For permissions, please e-mail: journals.permissions@oup.com.
Conflict of interest statement
Conflict of interest statement. T.C., H.Z., and X.L. have filed a patent application related to the work described in this manuscript. The other authors declare that they have no competing interests.
Figures

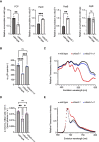
Comment in
-
Brown is the new green: Discovery of an algal enzyme for the final step of fucoxanthin biosynthesis.Plant Cell. 2023 Aug 2;35(8):2716-2717. doi: 10.1093/plcell/koad138. Plant Cell. 2023. PMID: 37195870 Free PMC article. No abstract available.
Similar articles
-
Green diatom mutants reveal an intricate biosynthetic pathway of fucoxanthin.Proc Natl Acad Sci U S A. 2022 Sep 20;119(38):e2203708119. doi: 10.1073/pnas.2203708119. Epub 2022 Sep 12. Proc Natl Acad Sci U S A. 2022. PMID: 36095219 Free PMC article.
-
Mediator subunit MED8 interacts with heat shock transcription factor HSF3 to promote fucoxanthin synthesis in the diatom Phaeodactylum tricornutum.New Phytol. 2024 Feb;241(4):1574-1591. doi: 10.1111/nph.19467. Epub 2023 Dec 8. New Phytol. 2024. PMID: 38062856
-
The light-harvesting antenna of the diatom Phaeodactylum tricornutum. Evidence for a diadinoxanthin-binding subcomplex.FEBS J. 2005 Sep;272(17):4339-48. doi: 10.1111/j.1742-4658.2005.04846.x. FEBS J. 2005. PMID: 16128804
-
Carotenoid biosynthesis in diatoms.Photosynth Res. 2010 Nov;106(1-2):89-102. doi: 10.1007/s11120-010-9589-x. Epub 2010 Aug 24. Photosynth Res. 2010. PMID: 20734232 Review.
-
Structural features of the diatom photosystem II-light-harvesting antenna complex.FEBS J. 2020 Jun;287(11):2191-2200. doi: 10.1111/febs.15183. Epub 2020 Jan 7. FEBS J. 2020. PMID: 31854056 Review.
Cited by
-
A key gene, violaxanthin de-epoxidase-like 1, enhances fucoxanthin accumulation in Phaeodactylum tricornutum.Biotechnol Biofuels Bioprod. 2024 Apr 2;17(1):49. doi: 10.1186/s13068-024-02496-3. Biotechnol Biofuels Bioprod. 2024. PMID: 38566219 Free PMC article.
-
The role of cis-zeatin in enhancing high-temperature resistance and fucoxanthin biosynthesis in Phaeodactylum tricornutum.Appl Environ Microbiol. 2024 Jun 18;90(6):e0206823. doi: 10.1128/aem.02068-23. Epub 2024 May 24. Appl Environ Microbiol. 2024. PMID: 38786362 Free PMC article.
-
Identifying the gene responsible for non-photochemical quenching reversal in Phaeodactylum tricornutum.Plant J. 2024 Dec;120(5):2113-2126. doi: 10.1111/tpj.17104. Epub 2024 Oct 30. Plant J. 2024. PMID: 39476224 Free PMC article.
-
Overexpression of PtVDL1 in Phaeodactylum tricornutum Increases Fucoxanthin Content under Red Light.J Microbiol Biotechnol. 2024 Jan 28;34(1):198-206. doi: 10.4014/jmb.2309.09018. Epub 2023 Oct 20. J Microbiol Biotechnol. 2024. PMID: 37957112 Free PMC article.
-
Distribution, Biosynthesis, and Function of Carotenoids in Oxygenic Phototrophic Algae.Mar Drugs. 2025 Jan 31;23(2):62. doi: 10.3390/md23020062. Mar Drugs. 2025. PMID: 39997186 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

