Early expression onset of tissue-specific effector genes during the specification process in sea urchin embryos
- PMID: 37101206
- PMCID: PMC10131483
- DOI: 10.1186/s13227-023-00210-2
Early expression onset of tissue-specific effector genes during the specification process in sea urchin embryos
Abstract
Background: In the course of animal developmental processes, various tissues are differentiated through complex interactions within the gene regulatory network. As a general concept, differentiation has been considered to be the endpoint of specification processes. Previous works followed this view and provided a genetic control scheme of differentiation in sea urchin embryos: early specification genes generate distinct regulatory territories in an embryo to express a small set of differentiation driver genes; these genes eventually stimulate the expression of tissue-specific effector genes, which provide biological identity to differentiated cells, in each region. However, some tissue-specific effector genes begin to be expressed in parallel with the expression onset of early specification genes, raising questions about the simplistic regulatory scheme of tissue-specific effector gene expression and the current concept of differentiation itself.
Results: Here, we examined the dynamics of effector gene expression patterns during sea urchin embryogenesis. Our transcriptome-based analysis indicated that many tissue-specific effector genes begin to be expressed and accumulated along with the advancing specification GRN in the distinct cell lineages of embryos. Moreover, we found that the expression of some of the tissue-specific effector genes commences before cell lineage segregation occurs.
Conclusions: Based on this finding, we propose that the expression onset of tissue-specific effector genes is controlled more dynamically than suggested in the previously proposed simplistic regulation scheme. Thus, we suggest that differentiation should be conceptualized as a seamless process of accumulation of effector expression along with the advancing specification GRN. This pattern of effector gene expression may have interesting implications for the evolution of novel cell types.
Keywords: Differentiation; Effector genes; Gene regulatory network; Sea urchin; Transcriptome.
© 2023. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests
Figures

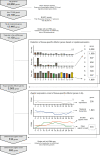





References
-
- Gilbert SF. Developmental biology. Massachusetts: Sinauer Associates Incorporated; 2013.
-
- Peter IS, Davidson EH. Genomic control process: development and evolution. Amsterdam: Elsevier Science; 2015.
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

